Home
> Musings: Main
> Archive
> Archive for January - April 2022 (this page)
| Introduction
| e-mail announcements
| Contact
Musings: January - April 2022 (archive)
Musings is an informal newsletter mainly highlighting recent science. It is intended as both fun and instructive. Items are posted a few times each week. See the Introduction, listed below, for more information.
If you got here from a search engine... Do a simple text search of this page to find your topic. Searches for a single word (or root) are most likely to work.
Introduction (separate page).
This page:
2022 (January - April)
April 27
April 20
April 13
April 6
March 30
March 23
March 16
March 9
March 2
February 23
February 16
February 9
February 2
January 26
January 19
January 13
January 7
Also see the complete listing of Musings pages, immediately below.
All pages:
Most recent posts
2026
2025
2024
2023:
January-April
May-December
2022:
January-April: this page, see detail above
May-August
September-December
2021:
January-April
May-August
September-December
2020:
January-April
May-August
September-December
2019:
January-April
May-August
September-December
2018:
January-April
May-August
September-December
2017:
January-April
May-August
September-December
2016:
January-April
May-August
September-December
2015:
January-April
May-August
September-December
2014:
January-April
May-August
September-December
2013:
January-April
May-August
September-December
2012:
January-April
May-August
September-December
2011:
January-April
May-August
September-December
2010:
January-June
July-December
2009
2008
Links to external sites will open in a new window.
Archive items may be edited, to condense them a bit or to update links. Some links may require a subscription for full access, but I try to provide at least one useful open source for most items.
Please let me know of any broken links you find -- on my Musings pages or any of my web pages. Personal reports are often the first way I find out about such a problem.
April 27, 2022
Briefly noted... Cataloging the outer solar system
April 27, 2022
A team of scientists has announced 458 previously unknown objects in the outer solar system, so-called trans-Neptunian objects (TNO). The discovery was "accidental". They were looking at the distant sky, but of course, their images included things in the way. So they developed computer analysis to extract novel closer objects. They published an earlier paper making a start, and now complete the study. Overall, what they found is about 20% of what is now known in the trans-Neptune part of the Solar System -- even though the current survey covered only a few percent of the region. Of particular note, they did not find Planet 9. More broadly, the results challenge current models for how the region formed.
* News story: A 6-Year Search of the Outer Solar System Turns up 461 new Objects (but no Planet 9). (Matt Williams, Universe Today, September 15, 2021. Now archived.) Links to an early version of the article, at ArXiv. (It also links to an earlier article on the project.) For the published article (which reports a slightly different number from what the title of this item suggests)...
* The article, which is open access: A Search of the Full Six Years of the Dark Energy Survey for Outer Solar System Objects. (Pedro H Bernardinelli et al, Astrophysical Journal Supplement Series 258:41, February 2022.)
* Background post about Planet 9: A ninth planet for the Solar System? (February 2, 2016).
Briefly noted... First in-human trial of an antibody against a prion disease
April 26, 2022
Prion diseases involve progressive neurodegeneration, and are inevitably fatal. No treatments have been shown to be useful. A new article reports a small trial of a monoclonal antibody against the prion, for Creutzfeldt-Jakob disease (CJD). Six patients were treated, with various doses for various times -- until they died or the supply of drug ran out. None showed a reversal of the disease, but three showed some slowing of the rate of progression. It was a tiny trial, and the results are limited. But even the hint of improvement means that the treatment warrants further testing and development.
* News story: Creutzfeldt-Jakob disease treatment shows promising early results. (Science Daily (University College London), March 17, 2022.) Links to the article, which is open access.
* Direct link to the article, which is open access: Prion protein monoclonal antibody (PRN100) therapy for Creutzfeldt-Jakob disease: evaluation of a first-in-human treatment programme. (Simon Mead et al, Lancet Neurology 21:342, April 2022.)
* "Comment" accompanying the article. It is not freely available, but it gives a good overview of the history of proposed treatments for prion diseases. Even the excerpt that is available may be useful, with some background. Investigating new treatments for Creutzfeldt-Jakob disease. (Inga Zerr, Lancet Neurology 21:299, April 2022.)
* This post is listed on my page Biotechnology in the News (BITN) - Prions (BSE, CJD, etc).
* For another approach to treatment... Treating prion disease with CHARM, to reduce production of the prion protein (August 21, 2024).
A special role for cyanide in the chemistry of early life?
April 25, 2022
Cyanide is best known as a poison. Suggestions that it might have played a role in the chemistry associated with the origin of life may seem surprising.
We should note that the poisonous role of cyanide is actually rather specific. It's bad for us, but that is because of a very specific biochemical step. There is no claim that cyanide is incompatible with life in general. And it may have been abundant.
Musings has noted an earlier article suggesting a role for cyanide in pre-biotic chemistry [link at the end]. That article suggested that cyanide could be a precursor to chemicals used for life.
We now have a recent article that suggests a more specific role for cyanide in the chemistry of early life. The key point is that, in certain situations, cyanide can effectively act as a reducing agent. That fills a need that has bothered chemists.
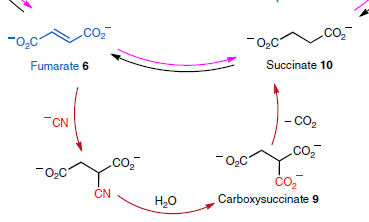
The following figure gives an example...
|
Reading "backwards"... Look at the reaction succinate (chemical 10) to fumarate (6). It is part of the TCA (or citric acid or Krebs) cycle, and is commonly at the bottom, which is why we see it "backwards". The reaction involves removing two H atoms to make the double bond found in fumarate.
The reverse reaction would be useful. In fact, many think that something like a reverse-TCA cycle was important in the chemistry of early life.
|
The lower loop of the figure shows how cyanide could play a role. There are three steps:
- The cyanide ion, -CN, adds to the double bond. (You can think of it as H-CN adding across the double bond.)
- The resulting nitrile is hydrolyzed. That produces a carboxylic acid, or (at neutral pH) a carboxylate ion, -CO2-.
- That makes a structure with two carboxylic acid groups very near each other. That is an unstable arrangement; one of the carboxyl groups is lost (decarboxylation). The result: succinate.
The black arrow at the top shows the direction of the TCA cycle; the pink arrow shows the direction of the reverse cycle. (Fragments of other arrows from the full figure are also visible at the top corners.)
This is the bottom part of Figure 1a from the article.
|
That loop transforms -- reduces -- fumarate to succinate, using cyanide. It effectively carries out the reverse of a step from the TCA cycle.
The authors also show a similar loop to reduce oxaloacetate to malate.
They not only showed the specific steps of the loops, but also looked at the implications for reversing the entire TCA cycle. The work pointed toward a shorter but useful variation of the TCA cycle, which could operate under a range of mild conditions (temperature and pH) without catalysts (even transition metal ions).
As always, remember that the work demonstrates some chemistry. It does not claim to show what actually happened in the real world origin of life on Earth. But the chemistry is novel -- and intriguing. It expands our "toolkit" of things that might have happened.
News stories:
* New role for cyanide in early Earth and search for extraterrestrial life. (Science Daily (Scripps Research Institute), February 3, 2022.)
* Cyanide and Life on Early Earth. (NASA Astrobiology, February 17, 2022.)
* News story accompanying the article: Prebiotic chemistry: Cyanide at the origin of metabolism -- The emergence of protometabolic reactions that evolved into today's metabolic pathways is unclear. Now, evidence suggests that the chemical origin of biological carbon metabolism may have relied on the versatility of a single primitive C1 feedstock molecule - hydrogen cyanide. (Saidul Islam, Nature Chemistry 14:123, February 2022.)
* The article: Cyanide as a primordial reductant enables a protometabolic reductive glyoxylate pathway. (Mahipal Yadav et al, Nature Chemistry 14:170, February 2022.)
Background post about cyanide in the chemistry of early life: Modeling the role of hydrogen cyanide in the pre-biotic formation of life's chemicals (September 8, 2019).
Other posts about pre-biotic chemistry include:
* Bitcoin and the origin of life (April 3, 2024).
* Miller-Urey revisited: the role of the glass container (January 22, 2022).
* The origin of life's chemicals: making an almost-Krebs-cycle in the lab (June 10, 2019).
Medical safety: What if pills were better labeled, so your phone could verify them?
April 23, 2022
How do you know a pill is authentic? Maybe you could have your phone check it.
Here is the idea, from a new article...
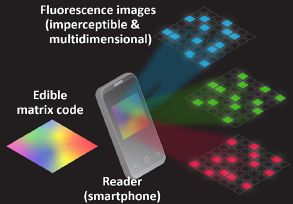
|
The pill would include a piece of code -- literally. It is actually a tiny bit of silk, modified to have a color pattern.
The phone images the piece of code, and sorts out the colors.
As the right side shows, there are three colors to analyze. Each has a 7x7 array of spots. Each spot can be on or off -- 0 or 1.
This is part of Figure 1 from the article.
|
That is 147 spots. There are 2147 possible codes. That is about 1044.
What is the purpose? The main purpose is to address the issue of counterfeit drugs, a significant and growing problem. The manufacturer would incorporate the code in a drug. The user's phone would read the pill code, and check that it is valid.
How is the coding done? As noted above, with silk. Silk is a protein. The scientists have made genetically modified silkworms that make variants of silk with regions from three fluorescent proteins (such as the familiar green fluorescent protein (GFP)). The rest is technology, to make tiny particles that have defined arrays of the color-producing proteins.
Each coding particle -- or label -- is about a millimeter across, and can be attached to an individual pill. It is barely visible -- a white speck. The colors appear only during the intense excitation used for fluorescence.
Is this safe? The only chemicals added by the labeling are the proteins, consisting of ordinary amino acids. It is possible for proteins to be toxic, but the proteins used here are familiar and generally regarded as safe. The authors suggest that the code pieces will be degraded by ordinary digestion, and they provide some evidence to support this.
The coding is done at the level of a batch. It is not about identifying individual pills (e.g., serial numbers), but types of pills. In terms of the main goal, the coding should authenticate the pills; it should contain information about the identity of the pill and the manufacturer. It could also include dosage, the specific batch (lot number), and the expiration date.
One can imagine how such coding could help a user keep track of their pills, reminding them to take the right pills, and such.
Liquid formulations can also be coded; the labeling is compatible with high concentrations of alcohol, found in some medicines. (One test example of a liquid was a whiskey -- another area where counterfeiting is a concern.)
Some current drug security measures are done at the level of the package. However, pills are commonly separated from the original package, for example by the pharmacy. The approach here does not depend on the original packaging. It also allows the consumer to check their own pills.
News story: Risks of Counterfeit Medications Are Rising: New Technology Could Spot Fakes With a Smartphone App. (SciTechDaily (from the journal publisher), April 13, 2022.)
The article, which is open access: Edible Matrix Code with Photogenic Silk Proteins. (Jung Woo Leem et al, ACS Central Science 5:513, May 25, 2022.)
Among other things you can do with a phone... Can you detect the SARS-2 virus with your phone? (March 2, 2021). Links to more.
For some background on GFP: Nobel prizes (October 8, 2008). Links to more.
Previous post about silk, from a spider in this case: A rigid silk basket (November 10, 2020).
Several Musings posts about silk are listed on my page Internet Resources for Organic and Biochemistry under Amino acids, proteins, genes.
My page Internet resources: Biology - Miscellaneous contains a section on Nutrition; food and drug safety.
April 20, 2022
Briefly noted... Exploring the mind of a jellyfish
April 20, 2022
Model systems have long played an important role in biology. Focus a lot of attention on studying one system in detail, then see how what we learned carries over to other organisms. Studying simple models can be good. For the "brain", studying jellyfish is about as simple as you can get. There is no discrete centralized brain. But there are neurons, and there is behavior -- surprisingly complex and coordinated behavior. A team of scientists has now begun to focus on the jellyfish Clytia hemisphaerica as a brain model. The organism is small (1 centimeter full grown), and transparent. In particular, the scientists have begun to develop genetic tools. For example, they can modify individual cells to produce green fluorescent protein.
* News story: How to read a jellyfish's mind. (Phys.org (Lori Dajose, Caltech), November 26, 2021.) Links to the article.
* Direct link to the article: A genetically tractable jellyfish model for systems and evolutionary neuroscience. (Brandon Weissbourd et al, Cell 184:5854, November 24, 2021.)
* Also recall... Briefly noted... Synapses in a sponge? (February 23, 2022).
* My page for Biotechnology in the News (BITN) -- Other topics includes a section on Brain. It includes a list of brain-related posts.
Separating isotopes of hydrogen using a MOF
April 18, 2022
MOF = metal-organic framework. It is now a broad term, for a range of materials. Musings has discussed some of them before, including in the context of gas separations [link at the end].
How good are they? Could they separate two gases that differ only in their isotopic composition? An experienced chemist knows there is a special case of that challenge, which is unusually favorable.
A new article reports using a MOF to separate hydrogen and deuterium gases, H2 and D2. The symbol D (= deuterium) is commonly used for the isotope of hydrogen with mass number 2, H-2.
The first figure lays the groundwork for the separation...
|
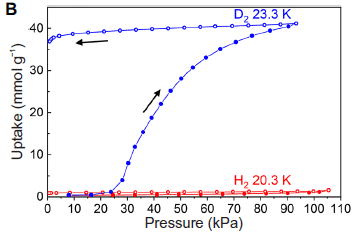
The two gases were tested (separately) to see how much bound to the MOF. The y-axis is the uptake of the gas into the MOF; it is plotted vs the gas pressure (x-axis).
The uptake of H2 is minimal.
The uptake of D2 is relatively high. It is also complex. The blue curve has two phases; note the arrows. The lower part of the blue curve shows that more and more D2 is taken up as P is increased. The upper part of the blue curve shows what happens as P is then decreased; the gas remains largely bound. We'll come back to the reason for this behavior later.
|
The two tests were done at different temperatures (T), as labeled on the figure. That may seem odd. Each test was done at the boiling point for that gas. Binding decreases at higher T; the uptake of H2 at the T used for D2 would be even lower.
This is Figure 1B from the article.
|
So the material takes up D2 much better than H2. What if we fed the MOF a mixture? The next figure shows such a test...
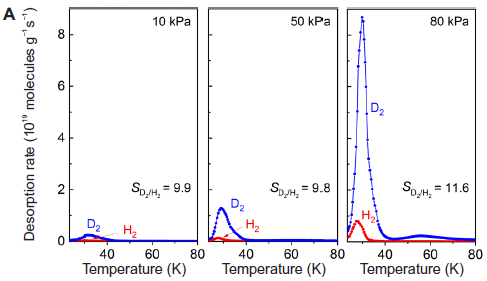
In each part, a 1:1 mixture of the two gases was fed to the MOF. The composition of the gas that came off the MOF (y-axis) was measured as a function of temperature (T; x-axis). (Actually, the y-axis shows the rate of desorption of each gas.)
The three parts are for various initial pressures. It will serve our purposes to focus on the right-hand graph, which is at the highest P -- and for which the graph is clearest.
You can see that much more D2 came off than H2. The authors calculate the selectivity, shown on the figure as SD2/H2, as about 12. (Similar values of S were obtained at lower P in the first two parts.)
This is Figure 3A from the article.
|
Looks promising. It may be possible to take a mixture of H2 and D2 gases, and effectively separate them by running them through a MOF. The authors think that what they developed here will be better (cheaper, less energy required) than current methods for separating these isotopes.
A little about the nature of the MOF and how things bind... This MOF is more complex than some we discussed earlier. The key here is that gas binding causes a structural change that opens up pores. The open pores then allow gas binding inside. In this case, D2 triggers pore opening, but H2 does not. Since the key step is opening the pores, once the gas is bound, simply lowering the pressure does not lead to gas release; go back and look at the complex behavior for D2 in the top graph, and the difference between raising and lowering P. The article discusses why the MOF transition is sensitive to the isotopes.
News story: Filtering deuterium with MOFs. (Nanowerk News (TU Dresden), April 14, 2022.) Includes a diagram showing the structural transition. The figure is a variation of Figure 1A from the article. (It is useful, but not entirely clear.)
The article, which is open access: Isotope-selective pore opening in a flexible metal-organic framework. (Linda Bondorf et al, Science Advances 8:eabn7035, April 13, 2022.)
Background post on MOFs: Cooperation: a key to separating gases? (March 28, 2014). Links to more.
A recent post about deuterium: Would weather forecasting be improved by considering the isotopes in atmospheric moisture? (May 2, 2021).
More hydrogen: Making hydrogen visible (July 2, 2022).
My page of Introductory Chemistry Internet resources includes a section on Nuclei; Isotopes; Atomic weights. It includes a list of Musings posts.
Making peptides in space: a new pathway
April 15, 2022
A team of scientists has proposed a new way to make protein -- and published it in an astronomy journal. Why an astronomy journal? It is actually a proposal for how peptides (small proteins) are made in space, from abundant materials on and around interstellar dust. It has been known that there are peptides out there, but there was no clear picture of how they were made.
The first part of the pathway is to make the monomers -- the subunits that are polymerized to make the peptide polymer. In conventional Earth-biology, the amino acids play this role. But outer space is not constrained by that 'rule'. Here is the proposed pathway to make monomers...
|
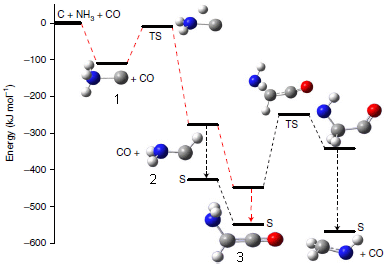
The main pathway starts at the upper left, with three chemicals considered available: C(gas), NH3, CO.
It proceeds downward through the three numbered compounds on the red-dashed pathway. The y-axis is an energy scale; going down means making chemicals that are more stable.
The first reaction is NH3 + C --> H3N-C (compound 1).
Then, one H moves from N to C, making H2N-CH (compound 2). (Between 1 and 2 is a higher-energy chemical, a transition state (TS).)
|
Compound 2 then adds CO, making H2N-CH=C=O (compound 3).
The C goes from having no bonds (in the original reactant, atomic C gas) to having 1, 2, and finally 4 bonds.)
The figure shows other possible reactions. But the authors suggest that the red-dashed pathway is most likely.
This is Figure 2 from the article. I have added numbers for three of the chemicals on the main (red) pathway.
|
Compound 3 is an amino ketene, not an amino acid. But amino ketenes, too, will get together and form amides. In fact, that is a spontaneous reaction, in contrast to the reaction of amino acids.
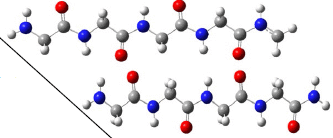
The following figure shows two of the polymers found in one lab experiment...
The two molecules shown here are the same, except at the right-hand end. If we ignore that point, the molecules are otherwise poly-glycine. Peptides with as many as 11 glycine units were found.
The H on the C of compound 3 is equivalent to the side chain H of glycine.
This is the inset of the top part of Figure 4 from the article. The figure shows mass spectra; these are the two chemicals being discussed there.
|
|
The authors provide theoretical calculations to show that the pathway is reasonable; see the top figure. And they provide some experimental data from lab work in support.
The claim is that this pathway could be how peptides, known to occur in space, are made there.
It is plausible that such peptides, made in space, were (or even are) delivered to Earth. Beyond that, we could speculate on why this might be of interest, but we will stop here, leaving the focus on the proposed pathway.
We should explicitly note that there is no claim that the proposed pathway, which may work in space, works (or ever worked) naturally on Earth.
News stories:
* How Life Came to Earth - Quantum Mechanical Tunneling Effect Might Play a Role. (SciTechDaily (Friedrich Schiller University Jena), February 13, 2022.)
* Peptides in outer space -- New chemical pathway allows for biomolecules to form on cosmic dust grains. (Max Planck Institute for Astronomy, Heidelberg, February 10, 2022.)
The article, which is open access: A pathway to peptides in space through the condensation of atomic carbon. (S A Krasnokutski et al, Nature Astronomy 6:381, March 2022.)
The work here is discussed, in part, in the context of origin-of-life chemistry. Among posts in that area:
* The role of tiny droplets in driving unfavorable reactions -- such as those needed to get life started (January 17, 2023).
* Encoding peptides from primordial RNA? (June 6, 2022).
* Was hydrogen the energy source for the origin of life? (February 1, 2022).
Also see: Quantum mechanical tunneling in DNA (July 27, 2022).
My page Internet Resources for Organic and Biochemistry has a section on Amino acids, proteins, genes. It includes a list of posts about making proteins.
April 13, 2022
Briefly noted... The gut microbiome and aging - in mice
April 13, 2022
It is known that the gut microbiome changes during aging. Whether that causes aging-associated changes has not been known. A recent article reports studies in mice, using fecal transplantation. Giving old mice the microbiome of young mice led to reduced aging symptoms. As always, we should be cautious about such microbiome studies, in mice or humans, but it is an interesting lead.
* News story: Poop transplant rejuvenates brain of old mice -- Replenishing the gut microbiome may reverse some aspects of cognitive decline due to old age. (Tibi Puiu, ZME Science, November 26, 2021.) Links to the article.
* My page for Biotechnology in the News (BITN) -- Other topics includes a section on Aging.
A solar cell that generates electricity at night
April 12, 2022
Reported in a new article.
Some data...
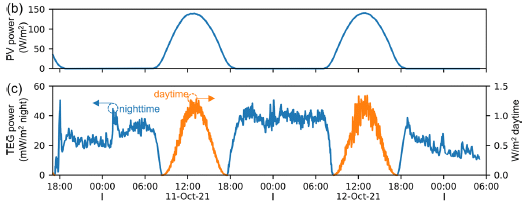
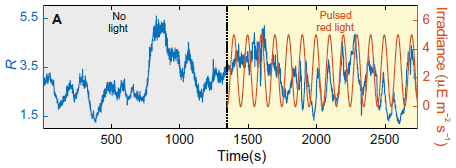
The figure shows the power produced by the parts of a modified solar cell system over the course of about two days.
The figure is complicated. For now, all we need to see is that power is generated most of the time, night and day.
We'll come back to the details after looking at the nature of the system.
This is part of Figure 3 from the article.
|
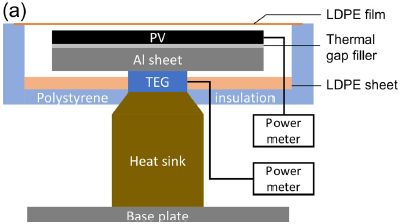
The next figure shows the design...
The main functional components:
- PV (photovoltaic), a traditional solar cell;
- a heat sink;
- between them, a TEG = thermoelectric generator.
There are separate power meters on the PV and TEG.
This is Figure 2a from the article.
|
Back to the top figure... Part b (top) shows the power from the PV cell. Part c (bottom) shows the power from the TEG. Also note that the TEG graph has two y-axis scales, one for the nighttime power (blue data; left-hand y-axis), and one for the daytime power (orange data; right-hand y-axis). (All powers are given per area of the PV cell.)
During the day, the Sun leads to direct production of electricity in the PV layer. That is traditional solar electricity. Further, the heat sink gets hot.
At night, the stored heat drives the thermoelectric generator to make electricity. Actually, the TEG is active during the day, too. In fact, it produces more power during the day than during the night. But it is the nighttime power that is of particular interest; this is solar power at night.
So, they have not cheated. They just stored some of the solar energy for later use. And when they do use it, they actually use a different process, not a solar cell per se.
Storing energy is not a new idea. We store water behind a dam, and use it when desired to run a generator. We could make solar electricity, and store extra electricity in a battery for later use; that deals with the limitation of making solar electricity only during the day.
What is novel here is how the solar energy is stored and used, by storing the heat and using it at night in a different generator. The idea is not entirely new, but this is perhaps the first demonstration that begins to look practical.
The authors suggest that their approach will be simpler and cheaper than the alternatives. Still. they see many ways to develop the process further.
News stories:
* Harvesting Energy at Night: Solar Cell Keeps Working Long After Sun Sets. (SciTechDaily (American Institute of Physics). April 5, 2022.)
* Photovoltaic cell harvests energy day and night. (Anne Fischer, pv magazine USA, April 6, 2022.)
The article, which may be freely available: Nighttime electric power generation at a density of 50 mW/m2 via radiative cooling of a photovoltaic cell. (Sid Assawaworrarit et al, Applied Physics Letters 120:143901, April 4, 2022.)
Among posts dealing with storing energy from intermittent sources...
* A better membrane for vanadium-based flow batteries (March 11, 2022).
* Storing energy from an intermittent source -- as compressed air under the sea (March 3, 2019).
* MOST: A novel device for storing solar energy (November 13, 2018).
Previous post about thermoelectric generators: A step toward a practical thermoelectric converter: get the oxide out (October 11, 2021).
Another post that is about both solar cells and TE devices: What to do with silicon waste from dead solar cells (May 23, 2022).
Posts about energy issues are listed on my page Internet Resources for Organic and Biochemistry under Energy resources.
Spider uses an external antenna to enhance hearing
April 9, 2022
Here is our subject, sitting on its acoustic antenna...
Larinioides sclopetarius, commonly known as the bridge spider. It is about a centimeter across. Bits of its web are visible.
This is the top figure in the news story from Cornell. It is credited there to one of the authors of the article.
|
The web may cover an area 10,000 times that of the spider.
The web consists of threads. Do the threads vibrate in response to sound waves? The following figure tests that...
|
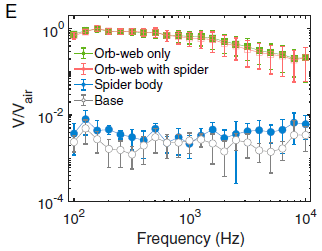
The figure shows the vibration of various things in response to an acoustic signal. The y-axis shows v/vair; that is the velocity of the target divided by the velocity of the air that was the stimulus. The x-axis is the frequency of the sound, from 100 to 10,000 Hz.
The top two curves are for the web, with or without a spider on it. The lower two curves are for the spider itself and for the base of the apparatus.
The web is the best receiver.
This is Figure 2E from the article.
|
That leaves one big question: Do the spiders actually use the web to receive sound signals? The article presents evidence to support that, but there is no easy way to show data. The idea is that the scientists direct auditory signals at the web, and show that the spider responds, for example, by turning toward the source.
Eardrums and threads detect sound differently, but they both work.
News stories:
* Orb-Weaving Spiders Use Their Web as Acoustic Antenna. (Sci-News.com, April 1, 2022.)
* Orb-weaver spider uses web to capture sounds. (Krishna Ramanujan, Cornell Chronicle, March 29, 2022.)
* Spiders use webs to extend their hearing. (Science Daily (Binghamton University), March 29, 2022.)
The article, which is open access: Outsourced hearing in an orb-weaving spider that uses its web as an auditory sensor. (Jian Zhou et al, PNAS 119:e2122789119, March 29, 2022.)
Movies. There are three short movie files with the article, under Supplementary Information. (10-65 seconds. Some sound, mainly the stimuli. Not critical.) These are somewhat interesting, but they are not labeled well enough to provide definitive evidence of the phenomenon.
Among posts about sensory responses in spiders:
* The ogre-faced spider, with massive eyes but no ears, detects the sound of prey using its legs (February 9, 2021). Leg hairs are another part of how spiders hear. But using the web leads to a big increase in sensitivity.
* Why the spider needs green light to find its food (March 3, 2012).
* How to seat a spider in front of the computer (September 28, 2010).
April 6, 2022
Briefly noted... Why some species of rockfish live for 200 years
April 6, 2022
It's a remarkable fact: species of rockfish vary in lifespan, from around 10 years to around 200 years. These species have diverged within the last ten million years. A recent article reports sequencing the genomes of 88 species of rockfish, thus allowing for comparison of the genomes. At this level, the work yields clues (correlations), not causal evidence. And most of the clues are the usual suspects. Keeping the immune system in check is important; inflammation is the enemy of aging. Good DNA repair is important. Species that live in deeper water tend to live longer; they have larger body size and smaller population size, common correlates of longer lifespan. At the basic science level, being able to compare the genomes of closely related organisms with quite different lifespans is an interesting development.
* News story: Pacific rockfish and the trade-offs of a long life -- Genome comparison of 88 rockfish species pinpoints genes associated with a long lifespan. (Science Daily (Robert Sanders, University of California - Berkeley), November 11, 2021.) Links to the article.
* News story accompanying the article: Aging: Long-lived fish in a big pond -- The genetic drivers of extreme longevity in Pacific Ocean rockfish are identified. (J Yuyang Lu et al, Science 374:824, November 12, 2021.)
* The article: Origins and evolution of extreme life span in Pacific Ocean rockfishes. (Sree Rohit Raj Kolora et al, Science 374:842, November 12, 2021.)
* My page for Biotechnology in the News (BITN) -- Other topics includes a section on Aging.
* There is more about genomes and sequencing on my page Biotechnology in the News (BITN) - DNA and the genome.
Another underground lake on Mars -- near the equator?
April 5, 2022
Scientists have recently reported what may be a large underground lake on Mars.
Here is the evidence -- and the map.
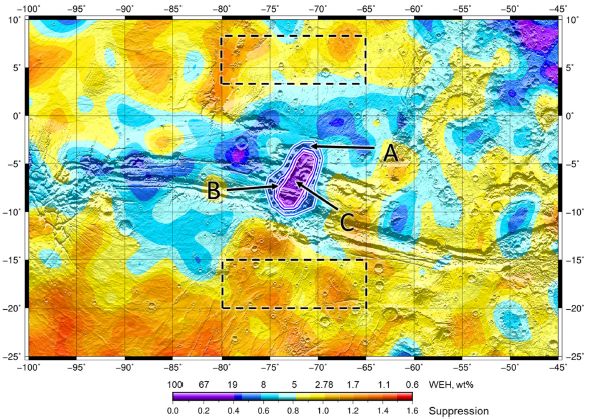
You probably found it, even before we described the map. But still...
There is a color key at the bottom. Look at the labeling called WEH. That stands for water equivalent of hydrogen -- a hint that things are not simple.
Briefly... Yellows and oranges are for dry regions (less than about 5% water). Blues are wetter. And purple is rather wet. There is one purple region, in a generally blue part of Mars.
The highest result for this purple region is about 40% water. That's wet, but not pure water. It could be slushy. And the water part is probably ice, if it is free at all.
The two regions shown with dashed boxes were used for reference measurements.
This is Figure 1 from the article.
|
What did they really measure? Hydrogen, as the label WEH suggests.
Actually, it is not hydrogen that they measured. They measured neutrons. The physics is complex, but here is a simple version... The soil is constantly bombarded with cosmic rays, which lead to neutron emissions. However, the observed neutron flux is affected by what atomic nuclei are in the soil. Hydrogen nuclei substantially suppress the neutron flux. The neutron flux is reported as "suppression", the lower scale on the key above. The suppression is relative to reference measurements. And the suppression is then interpreted as hydrogen, which in turn is interpreted as water.
The current work uses neutron measurements from a spacecraft in orbit at Mars, and thus provides data at high spatial resolution compared to previous work.
Hydrogen could be various things, but water is among the possibilities. High levels of hydrogen really do suggest water.
The error bars are huge. The highest value obtained for WEH, 40% at site C on the map above, comes with a +/- 1σ range of 16 to 100%, based on analysis of 13 pixels.
Where is this site? In a huge canyon, but not far from the surface (in the top meter). That makes it relatively accessible for on-site measurements.
Is there some concern that finding water in the top layer of soil on Mars, especially near the equator, is suspicious? Indeed. Perhaps it is tightly bound, but 40% water is a lot if it is tightly bound. Or perhaps the whole story is a misinterpretation of something.
We'll see.
News stories:
* ESA's Trace Gas Orbiter Finds Water in Martian Canyon System. (Sci-News.com, December 17, 2021.)
* Scientists find water in Mars' Grand Canyon. (Paul Scott Anderson, EarthSky, December 24, 2021.)
The article, which is open access: The evidence for unusually high hydrogen abundances in the central part of Valles Marineris on Mars. (I Mitrofanov et al, Icarus 374:114805, March 1, 2022.) The main part of the article is about four pages, and is very readable. That is followed by about four pages of Methods.
Posts about water on Mars include:
* Water on Mars? InSight finds none (September 19, 2022).
* A lake on Mars? (August 24, 2018). Links to more.
Previous post about Mars: Methane on Mars: a day-night effect (July 12, 2021).
next... How to make bricks on Mars (May 21, 2022).
Capturing carbon dioxide using gallium
April 4, 2022
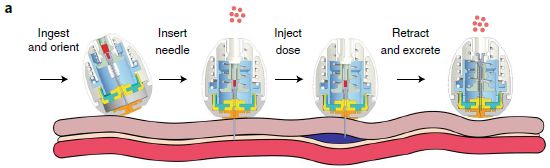
Here's the idea...
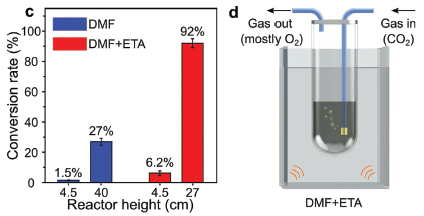
Start with part d (right); it shows the apparatus. The tube in the middle contains droplets of gallium metal suspended in a solvent. CO2 flows in, from the right; O2 flows out, at the left.
The waves at the bottom corners show that energy is being supplied -- by sonication. The fluid in the outer chamber is just to carry the energy.
Part c (left) shows some results. The y-axis shows the percentage of the CO2 converted. (We'll describe the reaction in a moment.) It's as high as 92% under the best conditions shown here. That includes the choice of solvent. DMF+ETA means dimethylformamide with 10% ethanolamine.
This is part of Figure 2 from the article.
|
Looks fairly simple, and it seems to work.
What's going on in there? The overall reaction is
CO2 (g) --> C (s) + O2 (g).
You saw the CO2 entering and the O2 leaving in part d above. The carbon is left in the liquid; it ends up floating to the top as thin sheets. The scientists think that the C could be a useful product.
The key step, of course, is getting started. The first step here seems to be
CO2 (dissolved gas) + Ga (l) --> Ga+ + CO2.-.
The superscript dot-minus on the CO2 product indicates that it is a radical anion; it has gained one electron, from the Ga. We'll skip the phases for the products, but they are probably near the metal surfaces.
That may be unfamiliar chemistry for many people, but it builds on things that are known. The radical anion is a common intermediate when reacting CO2.
The Ga reaction is a type common for metals, and has been studied for Ga.
In fact, such reactions have been studied for CO2 capture; they just don't work very well. A key development here is doing it with Ga, a liquid metal even under mild conditions. (The process is run at 40° C.) It is likely that using a liquid metal eliminates the problem of surface fouling, since there is not a fixed surface. The use of mechanical energy (sonication) to drive the process is also a novel development.
The system also contains solid nanoparticles of a gallium-silver alloy. Both the Ag chemistry and the solid surfaces are important.
The authors show that the process works; see the top figure here. They also show that the energy requirements are low. The details of the chemistry are not all clear, but this seems a useful start.
News story: Liquid metal proven to be cheap and efficient CO2 converter. (Neil Martin, University of New South Wales, October 13, 2021.)
The article, which may be "free to read": Liquid-Metal-Enabled Mechanical-Energy-Induced CO2 Conversion. (Junma Tang et al, Advanced Materials 34:2105789, January 6, 2022.)
Among posts on CO2 capture...
* Precipitating CO2 from air (July 30, 2022).
* Making carbon nanotubes from captured carbon dioxide (June 3, 2018). Links to more.
More about making use of the low melting point of gallium: Stretchable electric wires (January 22, 2013).
Carnivory in human ancestors: when did it start?
April 2, 2022
Perhaps you have heard that eating meat was an important development for humans.
Perhaps you have even heard that it started with Homo erectus, nearly two million years ago. Here are some data that might suggest that...
Caution... The article discussed here makes a complex statistical point. Don't try too hard to interpret the graphs shown here; they can only hint at the argument made in the article.
|
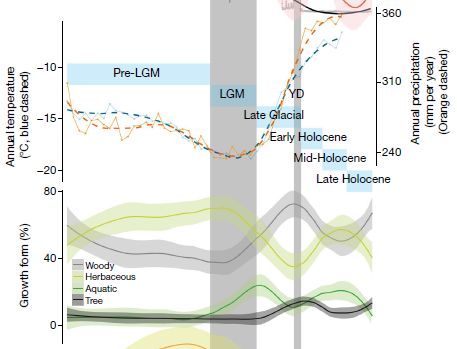
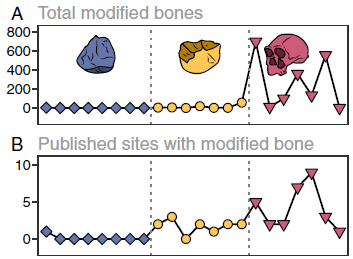
Part A (top) shows the number of "modified" animal bones found (y-axis) vs time (x-axis). Bone modifications such as cut marks are taken as an indicator that people were eating the animals. The time scale is from 3.4 to 1.2 million years ago; more about this below.
Plausibly, the graph shows an increase in modified bones in the right-hand part of the graph (that is, to the right of the right-hand dashed line; red triangles). That is about the time of Homo erectus.
Part B shows the number of published sites with modified bones over the same time span.
This is part of Figure 2 from the article.
|
The authors suggest that the increase in modified bones seen in part A might be an artifact of sampling. More sites from later times were sampled, so more modified bones were found at the later sites.
To test this, they made a model, assuming that there was not an increase in the real incidence of modified bones. How well does that model fit the data? The following figure shows a test of that...
|
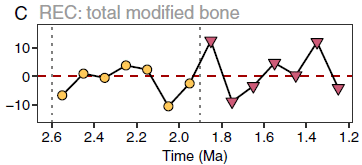
This graph shows the fit. The y-axis shows the "residuals", a measure of the discrepancy between the model and the data.
If there was a sustained increase in meat eating, there should be a consistent pattern of high residuals. That is, the values would be consistently above zero. They are not.
|
REC = residual evidence of carnivory.
The time scale here corresponds to the two right-hand segments of the first graphs, above (that is, yellow circles and red triangles). The x-axis of this graph is labeled (though I cut off the labeling for the top graphs).
This is part of Figure 3 from the article.
|
The authors conclude that there is no evidence for a transition to meat-eating over this time period, despite data such as the top figure above. That graph may be an artifact of having very limited data, biased toward later times.
It is also important to emphasize that the current article does not disprove a claim of increased meat-eating. The point is that the available data is not sufficient to allow any strong conclusion. That is a good lesson for studies of ancient events with limited data.
News stories:
* Human ancestor Homo erectus probably wasn't the carnivore we thought. (James Ashworth, Natural History Museum (London), January 24, 2022.)
* New Study Calls into Question the Importance of Meat Eating in Shaping Human Evolution -- Volume of evidence for meat eating can largely be explained by significant research attention on the time period after Homo erectus emergence. (GW Today, George Washington University, January 31, 2022.)
The article, which may be open access: No sustained increase in zooarchaeological evidence for carnivory after the appearance of Homo erectus. (W Andrew Barr et al, PNAS 119:e2115540119, January 24, 2022.)
More about the history of meat-eating in humans... Sliced meat: implications for size of human mouth and brain? (March 23, 2016).
More about the origin of carnivory: Venus flytrap: converting defense into offense (July 27, 2016).
Previous post mentioning H erectus, also with uncertainty: Evidence for early Homo floresiensis (July 5, 2016).
March 30, 2022
Briefly noted... Nanopore sequencing of proteins
March 30, 2022
Musings has discussed sequencing DNA by measuring changes in electrical current as an extended DNA chain traverses a narrow pore. A recent article extends nanopore sequencing to proteins. There are many technical issues to deal with. The main one here was developing a technique that facilitates multiple reads of the same protein molecule. It took many years for nanopore sequencing of DNA to go from being an intriguing idea to an established technique. The new work should be thought of as an early stage of extending it to protein sequencing. An early application might be more like identifying single-site variants of a particular protein.
* News story: Scanning a single protein, one amino acid at a time. (Nanowerk News (Delft University of Technology), November 4, 2021.) Links to the article.
* News story accompanying the article: Molecular biology: Toward single molecule proteomics -- Nanopore rereading of single proteins opens a pathway to next-generation proteomics. (Filip Bošković & Ulrich F Keyser, Science 374:1443, December 17, 2021.)
* The article: Multiple rereads of single proteins at single-amino acid resolution using nanopores. (Henry Brinkerhoff et al, Science 374:1509, December 17, 2021.)
* A background post: Nanopore sequencing of DNA using synthetic membranes (May 15, 2021).
Vitamin D, omega-3 fatty acids, and autoimmune disease?
March 29, 2022
A clinical trial. About nutritional supplements.
Do vitamin D or omega-3 (ω-3) fatty acids affect the incidence of autoimmune diseases in old people? (Fish oil is a common source for ω-3 fatty acids.)
It's fairly obvious what to do to test that. (We'll skip most of the details.) In fact, the trial piggy-backed on a trial of the effect of these supplements on cancer. People were getting the things anyway. May as well take multiple measurements.
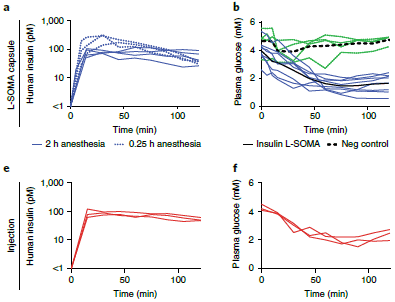
Here is a summary of the main results...
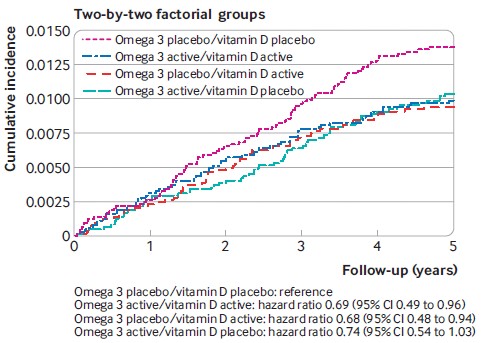
The graph shows the overall incidence of autoimmune diseases in four treatment groups, over five years of observation.
The four treatment groups: all possible combinations of +/- for each agent. Only one level of each agent was tested; it was rather high.
The big picture... One curve is high; the other three curves are all lower, and not too different from each other.
The high curve is for the all-placebo group. Those who received one or both of the supplements did better.
There are some statistics at the bottom. The reduction in hazard ratio is about 30%. The confidence intervals show that the results are barely significant; one tests as not significant (95% confidence interval includes a hazard ratio of 1).
The autoimmune diseases studied here include: rheumatoid arthritis, polymyalgia rheumatica,
autoimmune thyroid disease, psoriasis, and other.
This is part of Figure 1 from the article.
|
It is the best study yet of the effects of vitamin D and ω-3 fatty acids on autoimmune disease.
Is there reason to suspect there might be such effects? Sure. The supplements affect the immune system and inflammation. Anything more specific about how they work? Not much, especially considering the complexity of the system.
The results, as summarized above, are encouraging. But there are also limitations.
There were only about 26,000 people in the study, divided into four groups. Only 278 cases in total; the four sub-groups had 60-88 cases. (The graph shows that the incidence is about 1%.)
As noted, the trial used only one level of each supplement, a level that was rather high. Although both supplements are generally regarded as safe, there are limitations. Widespread use of them at high levels would raise questions.
The study lumps several autoimmune conditions. Would dissecting them be informative? (The article contains such information, but the already weak statistics become weaker.)
If these supplements reduce the incidence of autoimmune diseases, would they be useful in treating the people who do develop them? If so, that could be a more efficient use of the supplements. (The article notes that there have been some such trials -- small, and with results that were not particularly encouraging.)
An interesting article, which offers clues and raises questions. A typical nutrition trial.
Reminder... Musings does not give medical or nutritional advice. The post focuses on a single article; comments may even be intended to be provocative. (In this case, I find some of the commentary from the authors, away from the article, to be excessive.)
News stories:
* Vitamin D and Fish Oil Supplement May Reduce the Risk of Autoimmune Disease -- A study found that people who took vitamin D and omega-3 fatty acids had a lower rate of diseases like psoriasis and rheumatoid arthritis than people who took a placebo. (Becky Upham, Everyday Health, February 3, 2022.)
* Vitamin D supplements lower risk of autoimmune disease, researchers say. (Haley Bridger, Harvard Gazette, January 26, 2022.)
* "Opinion" piece accompanying the article; it may be freely available: Vitamin D and fish oil supplements and risk of autoimmune disease. (Karen H Costenbader, BMJ 376:o243, January 29, 2022.) From an author of the article.
* The article, which is open access: Vitamin D and marine omega 3 fatty acid supplementation and incident autoimmune disease: VITAL randomized controlled trial. (Jill Hahn et al, BMJ 376:e066452, January 29, 2022.)
More from the same trial: Do Vitamin D supplements prevent bone fractures? (August 8, 2022).
More about vitamin D:
* Briefly noted... Vitamin D and COVID-19? (June 23, 2021).
* Vitamin D: How much is too much? (July 9, 2013).
More about ω-3 fatty acids:
* Omega-3 fatty acids and the adaptation of fish to fresh water (August 3, 2019).
* Omega-3 fatty acids; fish oil (March 29, 2010),
My page Internet resources: Biology - Miscellaneous contains a section on Nutrition; Food safety.
An RNA replicator network
March 26, 2022
What would happen if there were only one kind of organism? Well, it would reproduce -- and mutate. Mutate to what? Well, to "everything", ultimately.
That's the idea. Is there evidence? Can we test such things, particularly for the very earliest stages in the existence of life?
A new article reports a test. Let's look...
The figure summarizes some key results. It is rather complex; we will just look at some highlights.
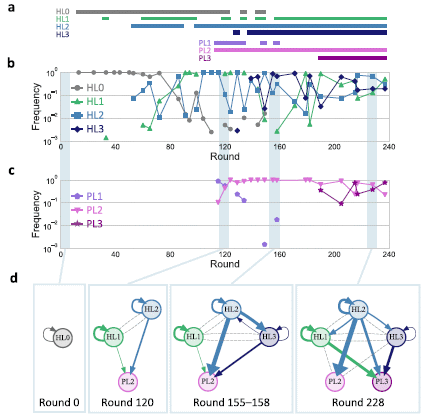
Start with part b. It shows the frequency of the original "organism" over time -- along with some others that arose during the test.
That original organism is called HL0; gray curve. You can see that its frequency starts at 100 (= 1); it is the only thing around. Its frequency varies after that -- and it is gone by round 160.
Three new organisms appear along the way, called HL 1 through 3. There is a bit of HL1 just before round 40 (see the isolated green triangle). It reappears at about round 60, and its frequency then steadily rises until it is the major strain. It fluctuates after that, along with the others.
We can see that the original organism (HL0) spun off new variant strains over the course of the test. Strains came and went; frequencies varied. Looks like the original organism was lost.
It all leads to lots of questions.
But first... The part b graph is confusing; let's look at a simpler way of summarizing the results.
Part a shows a single line for each strain, and simply shows whether it was or was not present (at the detection limit shown in part b). The information about frequency is lost in this form of the presentation, but it is useful. You can check frequencies if you want.
Part a, then, shows that HL0 was the only strain present at the start, and that it seems to have disappeared over time. Three other strains appeared at various times, and are the ones present at the end of this test.
What does it mean that a strain "disappeared"? It means that its frequency dropped below the detection limit. There is no claim that it has reached zero frequency. A strain that drops below the detection limit and does not reappear may have gone extinct, but the evidence here is not sufficient to show that. The strain may be present at a low level, even stably so.
This is Figure 3 from the article.
|
Let's step back at this point and ask, what is this "organism"? It is a small piece of RNA, which codes for its own replication enzyme (replicase). It is something like a virus, but even simpler; the scientists call it an RNA replicator. Doesn't a virus need a host cell to replicate in? Usually, but a friendly lab staff will do. The scientists take care of this RNA replicator, giving it the food and environment a cell would provide.
Part a shows more than the four HL strains. There are also some strains labeled with PL numbers. (The L is for "lineage".) These are parasites. They are distinct RNA species, but they are unable to grow on their own; they require a replicase from an HL strain.
Three major parasites are summarized in part a along with the HL strains. The frequencies for the parasites are shown in part c. That is just like part b for the HL strains, but each part has its own frequency scale. You can see that the parasites appeared relatively "late", but that two of these three parasites were still around at the end.
Finally, part d. It shows that the strains present at any given time were not growing independently, but were interacting. Any parasite strain was dependent on other strains to provide a functional replicase. Co-existing HL strains were also interacting. (Look only at the solid lines between strains. Dashed lines are for cases where no interaction was seen.)
Overall, then, we see that one kind of initial, very simple "organism" led to multiple and diverse "organisms", and to various interactions. It even seems that a system of some stability has been achieved. (Look at the frequencies of the five RNAs in parts b and c.) The scientists refer to this as a replicator network. It is more complex than a single type of molecule, but not yet an autonomous organism (no longer requiring quotation marks).
That's interesting.
Much of the discussion of the work is in the context of what might have happened in an early stage of life some billions of years ago. But we need to emphasize, the work does not show what did happen, but rather gives an example of what might have happened.
News stories:
* Scientists find new clues about the origins of life. (Andrei Ionescu, Earth.com, March 18, 2022.)
* Scientists Create RNA That Evolves on Its Own. This Could Be How Life on Earth Started. (Mike McRae, Science Alert, March 18, 2022.)
* New insight into the possible origins of life. (University of Tokyo, March 18, 2022.)
The article, which is open access: Evolutionary transition from a single RNA replicator to a multiple replicator network. (Ryo Mizuuchi et al, Nature Communications 13:1460, March 18, 2022.)
Also see...
* Encoding peptides from primordial RNA? (June 6, 2022).
* The magnesium dilemma: a step toward understanding how RNA might have been made in "protocells" (February 22, 2014).
* A novel type of polymer -- and its possible relevance to the origin of life (March 15, 2013).
March 23, 2022
Briefly noted... The Biden octopus
March 22, 2022

|
That's Syllipsimopodi bideni. It is about 328 million years old, and has ten arms. Soft-bodied octopuses usually don't fossilize well, so it is quite a find. It lends support to the notion that the 8-armed modern octopuses branched out from 10-armed ancestors (think squid). Why the Biden name? No reason is given, other than to honor the President.
|
* News story: Carboniferous Vampire-Squid-Like Cephalopod Had Ten Arms. (Enrico de Lazaro, Sci-News.com, March 9, 2022.) Includes an artist's conception of what the animal looked like, as well as a good view of the fossil. Links to the article, which is open access.
* The figure shown here is that "artist's conception", trimmed and reduced from the news story. There are actually two specimens in the figure, though the one at the lower right is hard to see.
* Previous such post... The Trump moth (January 31, 2017). Links to more.
* Next... Briefly noted... The Jerry Brown beetle (April 5, 2023).
* More about octopuses: Sleep stages in octopuses -- do they dream? (July 13, 2021). Includes an extensive list of posts about octopuses and other cephalopods.
* More about the vampire squid, which makes an appearance in this story: Quiz: What is it? (November 20, 2012).
|
Improving the accuracy of CRISPR: SuperFi Cas
March 21, 2022
A team of scientists at the University of Texas has made an improved version of the Cas protein, the enzyme part of the CRISPR gene editor. The new Cas is "more accurate"; it is called SuperFi Cas.
What do we mean by "more accurate"? Remember that Cas is guided to its target by a guide RNA, which base pairs with a DNA strand. As with any interaction between two nucleic acid strands, there is the possibility of mismatches occurring. Two strands with sequences that don't quite match pair up anyway. Such imperfect pairings can lead CRISPR to act at sites that were not intended: so-called "off-target" action.
The SuperFi Cas has a much lower frequency of such off-target actions.
Before we look at how the scientists figured it out, let's look at the result. The following figure summarizes results from the original Cas and the new SuperFi Cas...
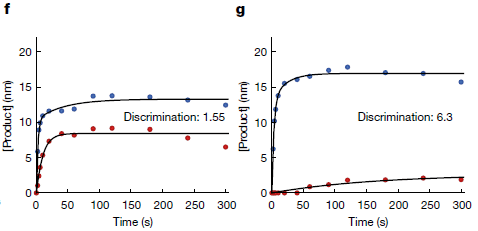
Part f (left) shows the time course of Cas action for the original Cas.
The test is done with an equal mixture of matched and mismatched DNA. The upper curve (blue points) shows the intended events. The bottom curve (red points) shows the off-target events.
Part g (right) shows the results of the same test for the SuperFi Cas.
You can see that the discrimination between target and off-target sites is much better for the new Cas. Further, the new Cas works fine on-target. That is, SuperFi Cas has reduced activity at the off-target site, but there is no performance penalty with the target site.
The difference between the two Cas proteins is even more dramatic for the initial rate. This point is discussed further in the article.
This is part of Figure 4 from the article.
|
How did the scientists get this improved Cas? They were studying the structure of Cas, specifically how it interacted with the DNA during the reaction. They realized that one piece of the protein generally seemed to do nothing -- unless Cas was bound to a mismatched DNA strand. It seemed that the role of this piece was to promote activation of Cas when bound to mismatched DNA.
To test that, they made a mutant version of the protein with that piece inactivated. The mutant behaved pretty much normally, except that it didn't activate Cas at mismatched DNA. That is, their hypothesis was right -- and they had made a Super Fidelity Cas.
This isn't the first mutant Cas with reduced activity at mismatches. However, it is the first such Cas that otherwise behaves normally. Making it required the understanding that came from studying the protein structure in detail during the process.
Does the mismatch action matter? For lab work, it is a nuisance; one screens the candidates from a CRISPR action to get the ones that are wanted. But if CRISPR is to be used in vivo, including in humans, errors are potentially serious.
One conclusion is that the normal Cas has a region that promotes its action with mismatched DNA. Does that make sense? Maybe. The natural role of the CRISPR system is to defend against invaders, such as viruses. Cas recognizes such things, and "kills" them. The mismatch enhancement seems to say, if in doubt, kill it. That may be a reasonable approach -- or outcome of natural selection.
We should note that the issue here is about one type of mismatch, and does not hold in general for all mismatches. The point is that it focuses on a type of mismatch where the natural Cas actually makes things worse (from our viewpoint).
News stories:
* Gene editing gets safer thanks to redesigned Cas9 protein. (Nanowerk News (University of Texas at Austin), March 2, 2022.)
* New Findings May Help Bolster CRISPR/Cas9 Accuracy. (Victoria Johnson, CGTlive, March 4, 2022.) Includes interview with one of the authors of the article, David Taylor.
The article, which is open access: Structural basis for mismatch surveillance by CRISPR-Cas9. (Jack P K Bravo et al, Nature 603:343, March 10, 2022.)
Posts about CRISPR are listed at: CRISPR: an overview (February 15, 2015).
Air filters that can kill
March 19, 2022
Air filters collect little things out of the air. Ordinary filters are not particularly effective at removing things as small as viral and microbial pathogens. And if they do remove them, they may end up serving as sources for what they collect. Wouldn't it be nice if air filters killed what they collect?
A recent article reports making such air filters. The approach is to simply bind disinfectant molecules to the filter material. It seems to work.
The first figure shows a lab test...
|
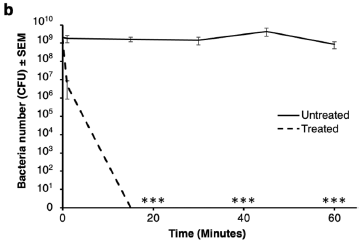
In this test. air filter material was exposed to a bacterial culture. One sample had been coated with a disinfectant. Bacterial survival (y-axis; log scale) was measured over time (x-axis).
The upper curve shows that the bacteria survived on the untreated filter material. However, they were quickly killed on the treated material.
The dashed line shows only one data point, with a substantial error bar, at 1 minute. All later values are "zero".
This is Figure 3b from the article.
|
That test was done with the bacterial pathogen commonly known as MRSA (methicillin-resistant Staphylococcus aureus). The full Figure 3 shows results for similar tests with two other microbes (E coli bacteria and the yeast Candida albicans); the results were about the same. Figure 4a shows results of a test with a virus (SARS-2, the COVID-19 virus); same idea, and same general result.
A biocide that gives 10-fold ("one log") killing is considered good. The results here far exceed that cut-off.
The next figure shows a test of durability...
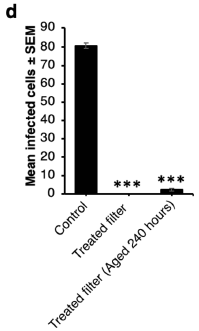
In this test, inactivation of the SARS-2 virus was measured with fresh filters and filters that had been subjected to a durability test for 240 hours.
The fresh filters were fully effective, and the aged filters were still very effective.
Note that the results are plotted in a different way for the virus work; fairly small virus numbers were used.
The authors consider the slight virus growth for the aged-filter sample to be not significant. It is not clear why they say that. (The results of a similar test with E coli are completely clean with the aged-filter sample. Figure 5c. Examination of the filters by electron microscopy did not suggest any damage. Figure 6.)
This is Figure 5d from the article.
|
|
The filters were then tested for three months in the real world: trains in the UK. Treated filters showed no recoverable microbes, as judged by colony-forming units (CFU). (No testing was done for viruses.) It is not clear what the sensitivity was, so we can't say what reduction there was, but obviously this is an encouraging result.
Overall, the article reports progress in making air filters that can kill pathogens. The process is compatible with current systems; it just requires a pre-treatment of the filters.
A company associated with the article is working on the commercial development.
And a reminder... Air quality is not an issue just for COVID. There are numerous respiratory diseases, including influenza.
What about HEPA filters? They do remove pathogens, but they are expensive, including the high energy cost for operation.
What is the disinfectant (biocide) here? Chlorhexidine digluconate. It acts mainly by disrupting membranes. It is effective against enveloped viruses, but not non-enveloped viruses.
News stories:
* CHX-treated air filters tested in trains found to kill SARS-CoV-2 and other microbes -- In both lab and real-world testing, chlorhexidine was an effective agent against many dangerous microbes that can circulate in public and health-care settings. (Amelia Williamson DeStefano, DentistryIQ, March 10, 2022.)
* New Antimicrobial Air Filters Tested on Trains Rapidly Kill Viruses - Including SARS-CoV-2/COVID-19. (SciTechDaily (University of Birmingham), March 10, 2022.)
The article, which is open access: Efficacy of antimicrobial and anti-viral coated air filters to prevent the spread of airborne pathogens. (Rowan Watson et al, Scientific Reports 12:2803, March 9, 2022.)
There has been less influenza during the COVID pandemic, presumably in part because of the measures introduced to reduce COVID transmission. Musings noted some early observations... How bad will the upcoming COVID-era winter flu season be? (September 25, 2020).
More about ventilation:
* Briefly noted... Improved ventilation to reduce COVID transmission (September 23, 2020).
* Indoor air pollution: is ventilation effective? (March 24, 2020).
My page for Biotechnology in the News (BITN) -- Other topics includes sections on Antibiotics (including disinfectants) and SARS, MERS (coronaviruses). They include lists of related Musings posts.
March 16, 2022
Briefly noted... The salt content of great 18th century violins from Cremona
March 16, 2022
What is the secret to the special sound of a violin made by Stradivari or other Italian masters of the era? It could be the chemicals used to treat the wood, according to a recent article. The original motivation might have been to preserve the wood, but the violin makers may have realized that the treatments affected the sound quality, and developed the treatments further. The actual work reported here involved analysis of small fragments of the instruments (left over from repair work). The general finding is that violins from a single family had consistent chemical characteristics. Then there is a lot of interpretation. It may seem that much is speculation, but the work leads to testable predictions -- and the possibility that instruments of Cremona quality could be made today.
* News story: Stradivari and Guarneri Treated Soundboards with Various Chemicals, Study Shows. (Sci-News.com, August 18, 2021.) Links to the article, which is open access.
* Also see: Better violins through better fungi? (March 4, 2013). Another suggestion, more or less the opposite of the current one.
* My page Internet resources: Miscellaneous includes a section on Art & Music. There is a list of related Musings posts.
Effect of water quality on the formation of microplastics
March 15, 2022
Plastic waste. It is all around us. It is obviously a mess, but there is much uncertainty about its forms -- and about how harmful the various forms may actually be.
One part of the story is that we make microplastics (MP) with ordinary use of plastic materials. Such "local" production of MP could well account for a significant fraction of our direct personal exposure to MP. As the name might hint, MP are the tiny particles most likely to get into our lungs.
A recent article makes an interesting point about our production of microplastics during ordinary use. Lab studies of such processes have generally used highly purified water, such as deionized water. The new work suggests that ordinary tap water leads to less production of MP. Why? The copper in the water forms a protective layer on common plastics, reducing their degradation.
The first figure is a simple test of the effect of copper ions in the water on the formation of microplastics from polypropylene (PP), one of the most common household plastics...
|
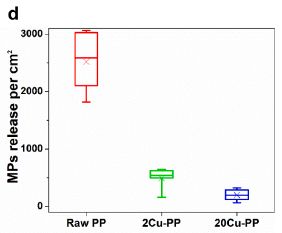
The results show the amount of MP released in a standardized test system.
The left-hand result is for the "raw" PP. The other two results are for PP pre-treated with two different levels of Cu2+ ions in boiling water. The lower level is typical of what is found in tap water.
You can see that the Cu2+ reduces the formation of microplastics substantially -- by about 80% in this test system.
This is Figure 2d from the article.
|
The following figure shows what the three materials look like...

|
The three materials are in the same order as above. In fact, the two figures should approximately line up.
|
You can see that the plastics treated with Cu2+ have become discolored. That is due to the formation of copper(II) oxide (CuO) on the plastic surface. The CuO serves as a protective layer.
This is Figure 2a from the article.
|
In another test, the scientists measured the release of microplastics from plastic kettles as a function of how many times water was boiled in them. The test used local tap water, with a Cu2+ content well below the relevant regulations (EU limit, 2 mg/L). Even a few boil cycles reduced the release of MP by more than 10-fold. After 2000 cycles, the release was reduced about 500-fold. (This test seems to lack a control, with purified water. But it is intended to mimic real-world use of the kettle.)
Other work in the article showed that Cu2+ similarly coats other common food-grade plastics. (Degradation tests were not done.)
The implications of the work are not entirely clear. One important message is that water quality matters; doing tests of MP production with highly purified water may be misleading. An interesting question that arises is whether products with plastic surfaces, such as kettles, should be manufactured with a protective coating.
News stories:
* Tap Water Produces Natural Protective Shield Against Harmful Microplastics. (AZoCleantech, October 22, 2021.)
* Minerals from tap water may be protecting us against microplastics -- Tap water may be doing us a big service. (Fermin Koop, ZME Science, October 22, 2021.)
The article, which is open access: Real-world natural passivation phenomena can limit microplastic generation in water. (Yunhong Shi et al, Chemical Engineering Journal 428:132466, January 15, 2022.) Passivation refers to the formation of a protective surface layer, which makes it passive.
More about microplastics: Consumption of microplastics by humans: the bottled water problem (September 20, 2019). Links to more.
Also see: History of plastic -- by the numbers (October 23, 2017). Added July 2, 2025.
A recent post about copper: Bacteria that make atomic copper (May 8, 2021).
Targeted degradation of the viral genome as a treatment for COVID?
March 13, 2022
Why not treat a viral infection by destroying the genome? Target a nuclease to the genome, and let it work.
In fact, a process for doing that, in principle, has been known for some time. It involves using siRNA -- small interfering RNA. siRNA is normally involved with regulating messenger RNA. Can we, instead, target it to the genome of an RNA virus? The idea is that the siRNA finds the genome, and recruits a nuclease. It has long seemed a good idea, but it has been hard to get it to work in practice. (The idea has similarities to how CRISPR works, but it is a distinct process.)
In a recent article, scientists reported trying to get siRNA to work against the SARS-2 virus, the agent of COVID-19. At least at lab level, they made progress.
The first figure shows the idea, for virus grown in cell culture in the lab...
|
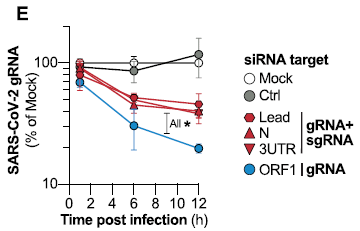
The experiment here reports the viral RNA made (y-axis) vs time during the infection (x-axis), for various conditions. The viral RNA was measured by the familiar PCR test. The results are expressed relative to a control.
The top two curves are for two types of control. One was a regular infection with no treatment. The other, called mock, used the siRNA procedure, but with no siRNA in it. Both gave similar results; the latter was set to 100% at each time point.
|
The other curves are for various types of siRNA. One of them is clearly best. It reduced the viral RNA by about 80% by 12 hours. This siRNA is targeted to a specific part of the viral RNA, called ORF1.
This is Figure 3E from the article.
|
That shows that siRNA can reduce viral RNA during an infection. Much of the effort involved trying several types; the results varied widely, as you can see above. RNA dynamics are complex during coronavirus infections. Some RNA sequences are abundant, and can sequester siRNA with minimal benefit; others may be inaccessible. The one that was best above was chosen for further work.
The next test used a more complex system: a piece of lung tissue.
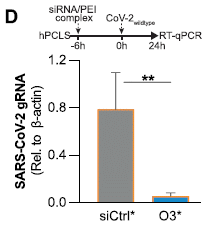
The top part shows the plan. The siRNA was given first. Six hours later, the tissue was infected. 24 hours after infection, viral RNA was measured. In this case, the viral RNA is shown relative to another cellular RNA (for actin).
Two types of siRNA were used. One was a control, which should not affect the infection. One was the O3 siRNA, the one against ORF1 as discussed above.
The results are clear: the O3 siRNA, targeted to the virus, reduced the viral RNA dramatically (by about 90%).
The siRNAs used here were modified to increase stability in the cell.
This is Figure 5D from the article.
|
|
It worked.
That is a prophylactic treatment: the drug was given before the virus. And, as noted, it is a lab-scale test, using a piece of lung tissue. So we might say that the work here establishes an approach, and suggests it is worth testing further.
News story: Targeted enzymes destroy virus RNA. (Nanowerk News (Technical University of Munich), March 2, 2022.)
The article, which is open access: Targeting genomic SARS-CoV-2 RNA with siRNAs allows efficient inhibition of viral replication and spread. (Shubhankar Ambike et al, Nucleic Acids Research 50:333, January 11, 2022.)
Another way to disrupt the viral genome: Could we treat COVID by driving it to an error catastrophe? (June 30, 2020).
More siRNA: Why does Zika virus affect brain development? (August 11, 2017). In this case, the siRNA was used to target a specific gene in lab work. The work had no implications for drug development.
More PCR: Ultra-fast PCR (March 14, 2023).
There is a BITN section for SARS, MERS (coronaviruses). It includes a list of Musings posts in the field.
A better membrane for vanadium-based flow batteries
March 11, 2022
Flow batteries separate the storage function of a battery from the electrodes. That allows a larger battery capacity to be serviced by the same electrodes. See the background posts listed below for an introduction.
One kind of flow battery is based on the rich redox chemistry of vanadium. But, as with flow batteries in general, practical implementation is lagging. For V-based flow batteries, one reason is that they "leak" -- lose charge. The common membranes used are too permeable to the vanadium ions.
A new article reports an improved membrane for V-based flow batteries.
The first figure shows the key development for the new membrane...
The idea is to make a material with tungsten trioxide (WO3) embedded between layers of graphene oxide (GO).
The stuff is called WO3@GO.
This is part of Figure 1a from the article.
|
The next figure shows how incorporating this new material into the membrane affects the permeability to VO2+ ions...
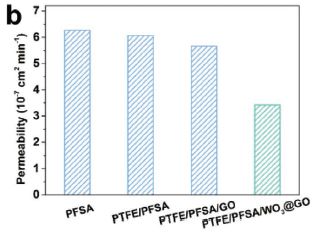
|
The bars show the permeability (y-axis) to VO2+ ions for four types of membrane.
At the left is the original membrane material. At the right is the new membrane, with the WO3@GO incorporated in it. The other two bars are controls, where some of the changes in the new membrane have been introduced, but not the WO3@GO.
You can see that the permeability of the new membrane is substantially reduced. The reduction is largely due to the WO3@GO.
|
One of the controls includes GO. But, without WO3 it is not nearly as good. The WO3 reduces aggregation of the GO. That was the original motivation for including it.
PFSA = perfluorinated sulfonic acid, also called by its trade name Nafion.
PTFE = polytetrafluoroethylene (Teflon).
This is Figure 3B from the article.
|
There are other effects of the new membrane. For example, the new membrane is actually better at conducting H+, which is good.
Does it matter? Is there an actual performance benefit, whatever the individual effects may be? The following figure shows one measure of that...
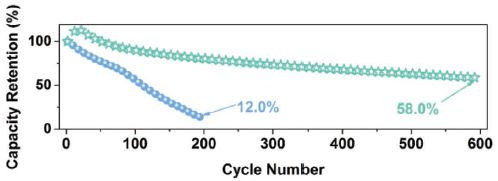
The graph shows how well the membrane ages. It plots capacity retention (y-axis) vs cycle number (x-axis). The lower (blue) curve is for the old membrane; the upper (green) curve is for the new membrane.
The new membrane ages much better than the old one. The old one is near dead at 200 cycles. The new one is still above half capacity at 600 cycles.
This is the bottom graph of Figure 5 from the article.
|
The authors note that the new membrane developments introduced here involve minimal additional cost, because the amounts of new materials used are small. The WO3@GO is at about 1%.
It is progress in developing practical flow batteries.
News stories:
* Novel ion exchange membrane improves performance of vanadium redox flow batteries. (Phys.org (Zhang Nannan, Chinese Academy of Sciences), November 11, 2021.)
* How Nanotechnology Can Improve Vanadium Flow Batteries. (Cvetelin Vasilev, AZoNano, December 17 2021.) Discusses current article and more.
The article: In Situ Grown Tungsten Trioxide Nanoparticles on Graphene Oxide Nanosheet to Regulate Ion Selectivity of Membrane for High Performance Vanadium Redox Flow Battery. (Jiaye Ye et al, Advanced Functional Materials 32:2109427, February 16, 2022.)
Background posts on flow batteries:
* A flow battery that uses polymers as the redox-active materials (January 8, 2016).
* Flow battery (January 4, 2016). Introduces flow batteries.
Other posts about batteries include... A battery made of paper, and activated by a drop of water (August 6, 2022).
More about energy storage: A solar cell that generates electricity at night (April 12, 2022).
Previous post mentioning vanadium: Upgrading ethanol? (April 11, 2016).
This post is listed for two sections of my page Introduction to Organic and Biochemistry -- Internet resources: Aromatic compounds and Energy resources. Both include lists of related Musings posts.
March 9, 2022
Briefly noted... A mutation that makes COVID worse protects against HIV
March 9, 2022
A year ago we noted an article identifying a genetic variant that increases the severity of COVID-19. Follow-up work showed that the frequency of the allele that leads to more severe COVID has increased over the last few thousand years. This might suggest that the allele has some beneficial effect. We now have a new article suggesting that the allele that leads to worse COVID protects against HIV. That is based on statistical analysis of three large databases (including the UK BioBank). How this works is not clear. The genetic region is involved with immune system function; it also happens to involve the receptor for HIV. It is not likely that the allele was originally selected for resistance to HIV, which is a modern virus. But it could have been selected based on resistance to some other virus, or it could have been selected for its effect on the immune system. Whatever the explanation, it seems to be an example of a genetic change that is good in one way and bad in another way.
* News story: Neanderthal Gene Increases Risk of Severe COVID-19 but May Offer Protection Against HIV. (Molly Campbell, Technology Networks, February 21, 2022.) Links to the article, which is open access.
* Background post: Susceptibility to severe COVID: role of a genetic region from Neandertals (February 20, 2021).
* Added June 11, 2025.
Next HIV post: Breaking latency of HIV, using a technology related to mRNA vaccines (June 11, 2025).
* There are BITN sections for HIV and SARS, MERS (coronaviruses). Each includes a list of Musings posts in the field.
Briefly noted... Electronic structure of lithium atoms
March 8, 2022
A lithium atom has only three electrons; one might think that the electronic structure of such a simple system would be well-established. But it is not, and that has implications for the use of lithium in batteries, which is so common. A recent article reports progress in establishing agreement between the theoretical and observed spectra for lithium metal surfaces, as well for lithium ions. It's quite technical, but this could turn out to be an important development.
* News story: Revealing lithium metal's electronic structure. (Nanowerk News (Lawrence Berkeley National Laboratory), January 26, 2022.) Links to the article.
* Previous post about lithium batteries: A lithium-ion battery that might be good for a million miles? (October 29, 2019).
* There is more about energy issues on my page Internet Resources for Organic and Biochemistry under Energy resources. It includes a list of some related Musings posts.
Clippane
March 7, 2022
Clippane is a new type of mechanically interlocked molecule.
Here is a diagram of one such molecule, reported in a recent article...

|
The clippane molecule consists of two pieces, shown in red and blue. (In this case, the red and blue pieces are the same.)
The red and blue pieces are interlocked -- mechanically. As a practical matter, the entire unit is one molecule.
This is the top part of Figure 3 from the article.
|
Here is the chemical structure of one of those pieces...
|
The R' groups (four of them, at the right) can be either hydrogen (H) or tertiary butyl (t-but or. more commonly, tBu). The latter is large, and restricts the motion of the ring structures. That is what leads to the effective interlocking.
The red part here corresponds to the loop part of the first figure. The blue parts here correspond to the bars of the first figure.
This is the upper right part of Scheme 2 from the article.
|
Much of the characterization of the product involves comparing the two forms. The one with R' = H has a free equilibrium between monomer and dimer forms. The one with the bulky R' = tBu does not; both the monomer and dimer can be isolated, and they do not interconvert.
Molecules with interlocking parts are not new. What is new here is that the parts are not rings. (There are rings in the chemical structure. However, the interlocking parts are not themselves rings.) Instead, the parts here look like tweezers.
The scientists had been studying these tweezer molecules for some time. The tweezers do tend to hold other molecules, as one might expect. And one tweezer molecule can hold another tweezer molecule. That observation led to the work reported here.
How do they make tweezer molecules? The flat arms and the linker (blue and red, respectively, of second figure) are brought together. The blue arms are interacting before the linker is connected. With the bulky R', connecting the linker traps some molecules in what will become the dimer form.
In the top figure, note that the red and blue pieces are perpendicular to each other. This is fairly clear in the middle. But also, the red and blue loops at the ends are perpendicular to each other. The article has figures that try to show the structure of the entire molecule; they are hard to follow.
News stories:
* New Class of Mechanically Interlocked Molecule. (Chemistry Views, November 1, 2021.) Good pictures.
* Welcome to the mechanically interlocked molecule family, clippanes! (Fernando Gomollón-Bel, Chemistry World, October 27, 2021.) Includes an animation showing the molecule during rotation; not sure how useful it is, but try it.
The article: Clippane: A Mechanically Interlocked Molecule (MIM) Based on Molecular Tweezers. (Susana Ibañez et al, Angewandte Chemie International Edition 61:e202112513, January 10, 2022.)
More interlocked molecules, of various types...
* Nanolympiadane: chemical synthesis of the Olympic rings (October 11, 2020).
* Molecular rings: little rings slip through big rings (July 17, 2018).
* Making lithium-ion batteries more elastic (October 10, 2017).
More tweezers... The smallest tweezers -- for pulling single molecules out of cells (March 30, 2019).
Flash photo-pyrolysis: converting banana peels to useful chemicals
March 5, 2022
Let's start with the bottom line. Here are some results from one run...
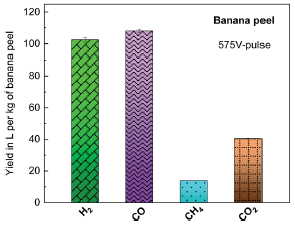
|
The exit gas stream contained large amounts of hydrogen and carbon monoxide. Useful feedstock for chemists; high energy content as a fuel. (The mixture is commonly called syngas.) Higher value than banana peels.
The results are expressed per kilogram of dried banana peel.
This is Figure 6 from the article.
|
How long did this take? About 5 microseconds.
That's not quite fair. What the scientists did was to subject the material to high energy pulses. The energy here is provided by flashes of light from a high-intensity xenon arc lamp. Each pulse is about a microsecond; pulses are delivered about every three milliseconds. Five pulses, that's about 5 µs -- over about 12 ms.
The light pulses heat the material to several hundred degrees, causing substantial degradation, with release of some gases. It is a type of pyrolysis process, where heat is used to degrade the material. This is a flash pyrolysis, stimulated by high energy light pulses. Importantly, pyrolysis is anaerobic. It generates reduced gases, suitable for further use. (Under aerobic conditions, such high temperatures lead to combustion -- to the oxidized products water and carbon dioxide.) The process can be done in ordinary steel vessels; high pressure is not involved.
The conversion is partial. Much is left as a porous residue, called biochar. The following figure shows the solids...
The top row shows before and after pictures, at the same magnification. Banana peel and biochar.
The bottom row shows two additional views of the biochar at higher magnifications; see the scale bars.
The main point with the photos is that the biochar is quite porous.
This is Figure 2 from the article.
|
H2 and CO are useful chemicals. Biochar may be useful. Banana peels are not. The process upgrades banana peels. The flash photo-pyrolysis process developed here leads to more of the gases, the preferred product, than previous work on gasification of biomass. And the biochar may even be better: highly porous and conductive.
This is lab-scale work; they are treating a 10 milligram sample. They tried various types of waste biomass, including corn cobs, orange peels, coffee beans and coconut shells in addition to the featured banana peels. There is little discussion of the economics, beyond noting that the process is a net producer of useful energy. They also note that the xenon lamps would not be practical at a large scale; the ideal might be to use solar light.
News stories:
* Getting hydrogen out of banana peels. (Nanowerk News (Ecole Polytechnique Fédérale de Lausanne), January 25, 2022.)
* Ultrafast pyrolysis makes hydrogen from banana peels. (Richard Thompson, Chemistry World, January 25, 2022.)
The article, which is open access: Banana split: biomass splitting with flash light irradiation. (Wanderson O Silva et al, Chemical Science 13:1774, February 14, 2022.)
More bananas and such: Food security: the potential of enhanced cultivation of enset (February 22, 2022). Links to more.
More syngas: A tower of (solar) power -- which makes kerosene (August 22, 2022).
More pyrolysis: Asteroids as a food source for astronauts (November 6, 2024).
Among posts about making use of biomass: Turning lignin into a useful product (April 11, 2015).
March 2, 2022
Briefly noted... Brain organoids grow eye precursor structures
March 2, 2022
Brain organoids developing in the lab from induced pluripotent stem cells under appropriate conditions may start to make visible eye structures. These optic cups include distinctive eye cell types, and are responsive to light.
* News stories:
- Lab-grown brain organoids develop primitive eyes. (Emma Green, BioNews, August 23, 2021.) Links to the article, and to a few other news stories.
- Lab-grown Brains Show Rudimentary Eye Development -- Stem cell-derived organoids treated with vitamin A spontaneously grew little optic cups. (Review of Optometry, August 19, 2021.)
* Previous post on brain organoids: Brain development: comparing human and chimpanzee lab-grown brain organoids (November 5, 2019).
* More... Human cortical organoids can survive -- and function -- in rat brains (October 22, 2022).
* My page for Biotechnology in the News (BITN) -- Other topics includes a section on Brain. It includes a list of brain-related posts.
* There is more about stem cells and related topics on my BITN page for Cloning and stem cells. It includes two extensive lists of related Musings posts.
A new form of carbon -- hard enough to scratch diamond
March 1, 2022
A picture, from a recent article reporting a new form of carbon...

|
The two photos show scratches on the surface of a diamond. The scratches were made by a piece of a new form of carbon.
The scale bar is probably 200 micrometers. (It has a label on it, but the label is unreadable.)
This is the right side of Figure 3C from the article.
|
The stuff is hard -- hard enough to scratch diamond. What is it? AM-III, one of the new forms of amorphous carbon reported in the article.
The following figure provides some data on its hardness, comparing it to other amorphous materials...
The first bar is for the various new amorphous carbon (AM) materials. AM-III is at the higher end of the range.
The new materials are several times harder than various other amorphous materials.
Diamond? It is not shown here, since it is not amorphous. But its hardness on this scale is about 60. (Diamonds vary -- and so do hardness scales.)
This is Figure 3B from the article. I have rotated it 90 degrees; the rotation makes it easier to read the labels for the bars. (The rotation also makes some of the other labeling look odd. For example, the B label is sideways and in the wrong corner.)
|
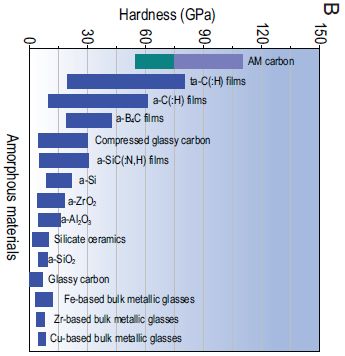
|
It's hard.
How do they make this stuff? About the same way one makes diamond from graphite in the lab: high pressure and high temperature. But instead of compressing graphite, they compress fullerene (C60; buckyballs). They tune the process, applying modest pressure over several hours. Depending on the details, they get a range of forms of amorphous C. AM-III is the hardest.
Might this stuff be useful? Well, it's hard. And conducts heat well. And is a semi-conductor. An interesting combination of properties.
Could it be made in useful amounts? That is open for now.
* Strongest glass in the world can scratch diamonds -- AM-III also works as a semiconductor, allowing it to transfer electrical current. (Tibi Puiu, ZME Science, August 10, 2021.)
* New Kind Of Glass That Is Harder Than Diamonds Created By Researchers. (Katie Spalding, IFLScience, August 11, 2021.)
* New ultrahard diamond glass synthesized -- It is the hardest known glass with the highest thermal conductivity among all glass materials. (Science Daily (Carnegie Institution for Science), November 24, 2021.)
The article, which is open access: Discovery of carbon-based strongest and hardest amorphous material. (Shuangshuang Zhang et al, National Science Review 9:nwab140, January 2022.)
"Carbon is one of the most fascinating elements due to its structurally diverse allotropic forms stemming from its bonding varieties (sp, sp2 and sp3). Exploring new forms of carbon has been the eternal theme of scientific research." From the beginning of the Abstract.
Posts on other new or unusual forms or behavior of C...
* A one-electron bond between two carbon atoms (October 23, 2024).
* Graphullerene (March 20, 2023).
* Briefly noted... A Möbius strip made of carbon atoms (September 7, 2022).
* Not graphene: a new flat form of carbon sheets (June 12, 2021).
* Making lonsdaleite -- with diamonds in it -- at room temperature (December 8, 2020).
* A new form of carbon: C18 (September 24, 2019).
Among posts about fullerenes... The smallest water bottle (January 5, 2011).
Posts about graphene, carbon nanotubes, and such (broadly, sp2-based C-materials) are listed on my page Introduction to Organic and Biochemistry -- Internet resources in the section on Aromatic compounds.
Added July 31, 2025.
A novel allotrope for another element: N6: A new form of nitrogen (July 31, 2025).
Does eating vegetables reduce the risk of cardiovascular disease?
February 28, 2022
Vegetables are good for you -- good for heart health. We just take that for granted. However, the evidence behind the statement is actually rather mixed.
The availability of large databases, such as the UK Biobank, allows tests of unprecedented size. A new article reports using the UK Biobank to test the effect of eating vegetables on death due to cardiovascular disease (CVD), and other causes.
Here is one of the summary figures...
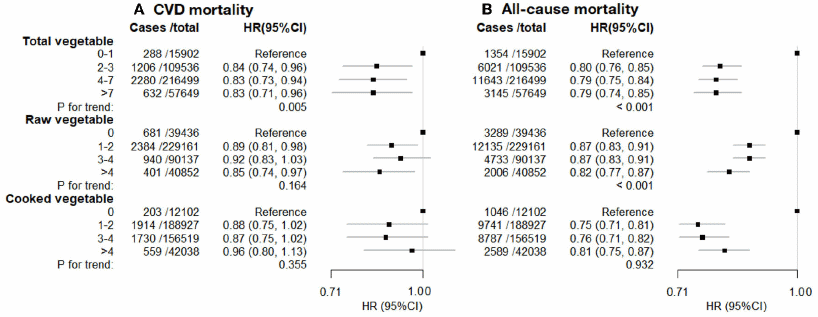
Let's start at the upper left, to see how the results are presented. This particular set is for the effect of total vegetables consumed vs deaths from CVD.
Vegetable consumption is presented (extreme left column) as number of heaped tablespoons per day, from 0-1 to >7. The first group (0-1) is taken as the reference; the other groups are compared to this reference group.
What is calculated is the hazard ratio (HR): the risk of death from CVD for a given level compared to the reference level. The HR is tabulated and also shown pictorially, with 95% confidence limits. The reference case is set to 1.
You can see that the other levels of total vegetables all give HR about 0.85 -- about 15% lower than the reference level. There is no evidence for a trend with amount of vegetables.
That upper left set is for the effect of total vegetables on deaths from CVD. The two sets below that are for raw and cooked vegetables, respectively. Part A (left) is for CVD deaths; part B (right) is for total (all-cause) deaths.
Caution... The horizontal scaling for the graphs is different in the two parts. You can make visual comparisons vertically (within a part). But if you want to compare parts A and B, check the numbers.
This is Figure 3 from the article.
|
What does it mean? It is not obvious, despite following 400,000 people.
We noted that for the upper left case, all higher levels of vegetable consumption were about the same: a somewhat reduced HR for CVD death, but no dose effect. In fact, that substantially holds over the entire figure. Of the 18 HR values shown, 16 are in the range 0.79-0.89. That range is narrower than the confidence limits in part A, and similar to those in part B (which are narrower because there are more deaths for Part B). Most of the values seem significantly lower than the reference value, but there is little evidence for any dose effect.
What may be most important is what is not shown here. The results shown above are from complex statistical analysis. One important part of the analysis is accounting for confounding variables. The authors discuss this; they found that including more possible confounding variables reduced the apparent effect size. They are concerned that better analysis of the confounding variables would reduce it further. That is, they are concerned that the analysis for confounding variables is just not right.
There is nothing here that suggests vegetables are bad for you. The authors do not intend to discourage eating vegetables.
The point is more that the work shows how difficult it is to analyze human nutrition, even with vast databases available.
Further comments...
The following statement from the article hints at the problem: "Participants with higher levels of total vegetable intake were more likely to be women, better educated, and residents of an affluent area, with lower mean BMI and higher levels of physical activity, and less likely to be smokers." (page 3, bottom) The question is whether confounding factors such as those are properly taken into account in the analysis. The authors suspect they are not.
The table shows results for raw vs cooked vegetables. Indeed, a primary goal of the work was to distinguish the effects of raw vs cooked vegetables. However, after seeing the results, such as those above, any conclusions on that issue are questionable.
Both the Introduction and Discussion of the article talk about related studies. The big message, as the authors note, is the inconsistency between studies.
News stories:
* Eating vegetables does not protect against cardiovascular disease, finds large-scale study -- Previous positive studies may not have sufficiently corrected for confounding socio-economic and lifestyle factors, suggests new analysis . (EurekAlert! (from the journal), February 21, 2022.)
* Vegetables: Our Cardiovascular Besties, or Just Another Acquaintance? (Chuck Dinerstein, American Council on Science and Health, February 21, 2022.)
* Expert reaction to study looking at reported vegetable consumption and risk of cardiovascular disease . (Science Media Centre, February 21, 2022.) A number of experts -- quite a large number in this case -- offer diverse evaluations of the work. If you are seriously interested in the topic, read this item.
The article, which is open access: Raw and Cooked Vegetable Consumption and Risk of Cardiovascular Disease: A Study of 400,000 Adults in UK Biobank. (Qi Feng et al, Frontiers in Nutrition 9:831470, February 2022.)
More about diet and heart disease: The "paleo diet" -- a trial (August 11, 2019). Links to more.
My page Internet resources: Biology - Miscellaneous contains a section on Nutrition; Food safety. It includes a list of related Musings posts.
Do mosquitoes remember encounters with sub-lethal levels of pesticides?
February 26, 2022
Insecticides are useful against mosquitoes. However, they lose effectiveness over time. Mosquitoes become resistant to them.
Is it possible that the mosquitoes also learn to avoid insecticides? A new article addresses the question.
The following figure shows some of the results...
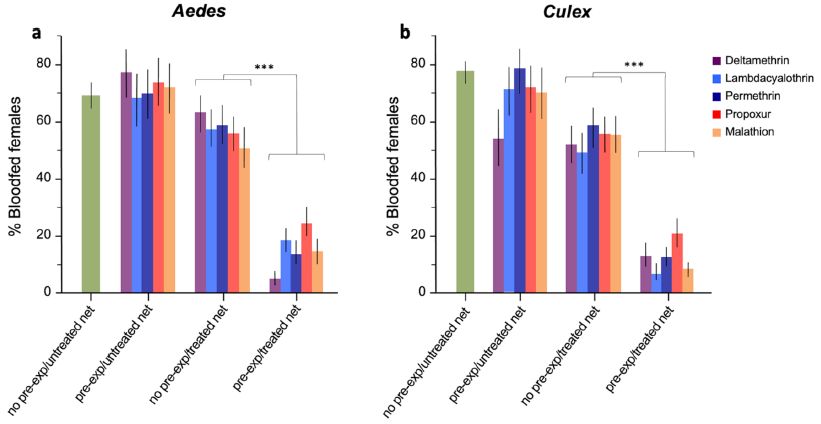
In this test the frequency of (female) mosquitoes that went on to feed (y-axis) was tested under various conditions. They had to cross a net, which was or was not treated with an insecticide. The other key variable was whether or not the mosquitoes had been pre-exposed to the insecticide.
Five insecticides were tested; their results are shown with different colors, as listed in the key at the upper right. Since the results are broadly the same for all the insecticides, we'll just discuss them as a group.
Part a (left) shows the results for Aedes mosquitoes.
There are four data sets; the results for the right-hand set of bars are strikingly low. That is for mosquitoes that have been "pre-exp"[osed] to the insecticide encountering a net "treated" with that insecticide.
The two data sets before that are for pre-exposed mosquitoes encountering an untreated net (without the insecticide), or mosquitoes without pre-exposure encountering a treated net. Meeting only one of those two conditions did not interfere with feeding. The first data set (represented by only one bar. since there is no insecticide for this condition) is the control with neither condition: no pre-exposure and no net treatment.
The pre-exposure killed about 30% of the mosquitoes. That level was chosen to try to get a condition where it was clear that the survivors had been significantly exposed, but were not resistant mutants. The testing was done 24 hours later.
This is Figure 1 from the article.
|
The conclusion is clear: pre-exposure to an insecticide leads to avoidance.
Part b (right side) shows the results of testing Culex mosquitoes. The pattern is the same.
Other results showed that pre-exposure to the insecticide increased the survival of the mosquitoes when exposed to the treated net, because they avoided it.
The finding that mosquitoes have this associative learning is not entirely new, though it is usually not considered in developing mosquito abatement practices. The point of the article is that it uses standardized assays to test for the learned behavior.
News story: Biology: Mosquitos learn to avoid pesticides after single non-lethal exposure. (EurekAlert! (from the journal), February 17, 2022.)
The article, which is open access: Standardised bioassays reveal that mosquitoes learn to avoid compounds used in chemical vector control after a single sub-lethal exposure. (Seynabou Sougoufara et al, Scientific Reports 12:2206, February 17, 2022.)
A recent post about developing a novel insecticide against mosquitoes: A trap to attract -- and kill -- mosquitoes (October 26, 2021).
Posts on the general nature of mosquitoes are included in the listing at a section of my page Biotechnology in the News (BITN) -- Other topics on Malaria.
More about pesticides: The role of ants in agricultural pest control (September 12, 2022).
February 23, 2022
Briefly noted... Synapses in a sponge?
February 23, 2022
Well, no. But they do have genes for proteins considered characteristic of synapses. A recent article showed that these genes are expressed in certain cell types associated with digestion. The role of the synapse-like proteins is not clear, but perhaps they are involved in coordinating digestive function between cells. Is this role of synapse genes in sponges a step in the development of nervous systems? Again, that is not clear at this point. It is an intriguing article.
* News story: Sponge cells hint at origins of nervous system -- Synapse genes help cells to communicate in sponge's digestive chambers. (Max Kozlov, Nature, November 5, 2021.) Good overview, including the implications. Links to the article.
* A post about a sponge -- one with an unusual eating habit: Quiz: What is it? (October 31, 2012).
* A post about the nervous system of a simple animal, with some discussion of the origins of nervous systems: A novel nervous system? (July 20, 2014).
* Also... Briefly noted... Exploring the mind of a jellyfish (April 20, 2022).
* My page for Biotechnology in the News (BITN) -- Other topics includes a section on Brain. It includes a list of brain-related posts.
Food security: the potential of enhanced cultivation of enset
February 22, 2022
Let's start with a question. What kind of banana, or banana-like plant, has gotten recent attention as a food crop that should be grown on a much larger scale? You can answer true or false.
The answer is false -- the false-banana, more properly known as the enset, Ensete ventricosum.
Enset is cultivated in a limited area of the highlands of Ethiopia. But biologists at the Royal Botanic Gardens (Kew) realized that its properties should allow it to be grown over a much wider area, even with projected climate change.
The following map summarizes some of their findings from modeling...
The map shows a region of east and southern Africa.
The striped region shows the "current range" of cultivation of enset. (It is largely just to the right of the lower part of the key.)
The green and red regions show where the scientists think it could usefully be grown. Darker colors are for regions they consider higher priority. (The specific map here is for one particular projected scenario for 2070.)
Green is for the current domesticated enset. Red is for wild enset. (CWR = crop wild diversity.) The wild strains are not edible, but represent diversity that could be developed.
Numbers along the border of the figure are degrees of latitude and longitude. The width of the figure is about 1500 km.
This is Figure 6b from the article.
|
The analysis suggests that the plant has the potential to serve several times more people in the region than it does currently.
Why isn't it grown over a wider area now? There may be multiple factors, but the scientists suggest that one reason is that the current region of domesticated enset is rather isolated, limiting its spread.
The article includes discussion of socio-economic and political issues that affect distribution of crops, as well as the biology and climate factors.
The work is an example of looking at the potential of crops that are generally not well known. The African Orphan Crop Consortium alone has identified 101 crops worthy of consideration.
News stories:
* 'Sustainable, nutritious and unique' Ethiopian orphan crop promises food security solution. (Oliver Morrison, foodnavigator.com, January 27, 2022.)
* The 'false banana' feeding millions -- Could a remarkable banana relative feed over 100 million people? (James S Borrell & Olef Koch, Royal Botanic Gardens (Kew), January 21, 2022.) From two of the authors of the article.
The article, which is open access: Modelling potential range expansion of an underutilised food security crop in Sub-Saharan Africa. (O Koch et al, Environmental Research Letters 17:014022, January 2022.)
Previous posts on false bananas: none.
Posts on true bananas include:
* Male mice are stressed by the odor of bananas or pregnant females (June 14, 2022).
* Flash photo-pyrolysis: converting banana peels to useful chemicals (March 5, 2022).
* Briefly noted... Using CRISPR to make better bananas (August 25, 2021).
* Measuring radiation: The banana standard (April 17, 2011). Links to more.
Posts on broader agricultural issues include... Evaluating the sustainability of food systems (December 5, 2022).
My page Internet resources: Biology - Miscellaneous contains a section on Nutrition; Food safety. It includes a list of related Musings posts, including some on food security.
Could ships harvest plastic from the ocean -- and use it for fuel?
February 21, 2022
Plastic waste is accumulating. One famous accumulation is the Great Pacific Garbage Patch (GPGP). And ships are working on cleaning it up, harvesting the plastic and bringing it in. It's a slow and expensive operation. Can we do better?
A recent article explores the possibility that the clean-up could pay for itself, or at least fuel itself. The idea is to convert the plastic to fuel on the ship, and use the fuel for the operation.
Will it work? Maybe. Let's look at a couple pieces of the analysis...
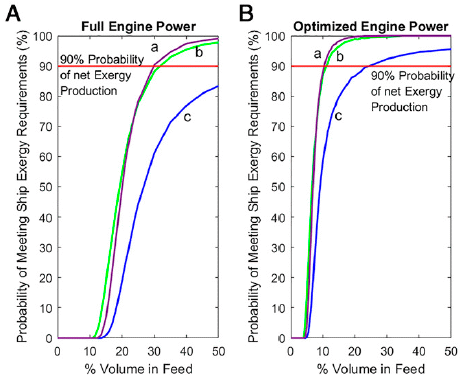
This figure examines how much plastic would be needed to make the process self-sufficient.
The x-axis is the percent of plastic in the feed. The y-axis is the probability that the process would be self-sufficient. (That the results are given as probabilities reflects that there are many uncertainties in the analysis. which is done by computer simulation.) The various curves are for various details.
The two parts make different assumptions about how the ship operates. Part A is for the ship at full engine power; part B is for the ship operating in an energy-optimized manner.
The three curves in each part make different assumptions about the nature of the plastic. Curve a is perhaps of most interest; it is for a 2:1 mix of polyethylene and polypropylene, considered typical of ocean plastic.
Looking at the 'a' curves... Part A says that about a 30% concentration of plastic would be needed; Part B says that about a 10% concentration of plastic would be needed. Ship efficiency matters.
This is Figure 2 from the article.
|
An optimistic interpretation is that the process might work if the feed were at least 10% plastic.
That's a lot. You may have the impression that there is a lot of plastic out there, but the ocean is not 10% plastic.
Practical processes use booms to collect the plastic, delivering it to the ship in a relatively concentrated form. It takes many booms deployed around the ship to get the feed to a useful level.
The following figure brings together several factors...
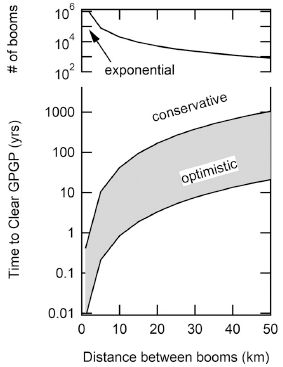
|
The main (bottom) part of the figure shows how long it would take to clear the GPGP (y-axis) as a function of the distance between booms (x-axis).
Note that the time scale is in years, on a log scale. 0.01 year is about 4 days. The top of the graph is a millennium.
That is quite a time range. That is the point. But a clean-up time on the order of 1 to 10 years may be possible.
The upper part of the graph shows how many booms would be used (y-axis) vs the spacing (the original x-axis).
This is Figure 4 from the article.
|
There are many uncertainties, as you can see. The point of the current article is to put the proposal on the table for more detailed consideration. It seems that it might work for large concentrations of ocean plastic, such as the GPGP.
Further comments...
The main process considered here for converting plastic to useful fuel is hydrothermal liquefaction. Other processes could be considered.
One consideration is that the clean-up must be faster than the degradation to smaller pieces, which become harder to harvest.
An important benefit of the proposed approach is greatly reduced fuel needs for the ships. With current processes, the ships collect the plastic, and then transport them back to port. With the proposed process, the travel just to transport plastic would be eliminated. The recovered plastic would fuel the ship and the conversion process.
News stories:
* How about powering ocean cleanup ships with the plastic they collect. (Prachi Patel, Anthropocene, November 25, 2021.)
* Could we use plastic to end the ocean's plastic problem? A new paper says: "Yes!". (Alexandru Micu, ZME Science, November 2, 2021.)
* Ocean cleanup vessels may burn plastic pollution as 'blue diesel'. (Environmental Science & Engineering Magazine, November 8, 2021.)
The article, which is open access: Thermodynamic feasibility of shipboard conversion of marine plastics to blue diesel for self-powered ocean cleanup. (Elizabeth R Belden et al, PNAS 118:e2107250118, November 16, 2021.)
Among posts about the plastics problem...
* What's in the North Pacific Garbage Patch? (September 20, 2022).
* Turning waste plastic into fuel -- a solar-driven process? (October 9, 2018).
* History of plastic -- by the numbers (October 23, 2017). Links to more.
There is more about energy issues on my page Internet Resources for Organic and Biochemistry under Energy resources. It includes a list of some related Musings posts.
Tau and ALS
February 19, 2022
The brain protein tau is involved in a variety of neurodegenerative diseases, collectively called tauopathies. Alzheimer's disease is perhaps the one that gets the most attention.
There is evidence implicating tau in amyotrophic lateral sclerosis (ALS). A new article provides more evidence, and even offers a clue for treatment.
Here is one graph from the new article...
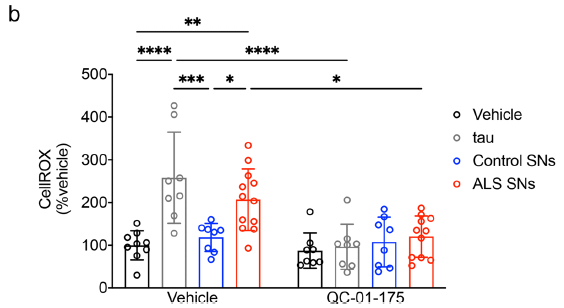
This work involves measuring the reactive oxygen species (ROS; CellROX, on the axis label, is a chemical used to test for ROS) in lab-grown cell cultures under various conditions. The ROS are thought to be involved in the development of ALS.
Caution... The term 'vehicle' is used here to mean two different things. If you realize that, the meanings are clear from context; in each case, the 'vehicle' is a control for what is being tested at that point.
The first four bars (to the left, the 'vehicle' set) lay the groundwork. The left-hand bar is the control for this set (and is labeled 'vehicle' in the key at the far right). The level of ROS for this case was set to 100%.
The second bar shows what happened if tau was added; the level of ROS doubled. The final two bars of the set are for the addition of material called synaptoneurosomes (SN), either from control tissue (blue) or ALS tissue (red). The SNs from control tissue had little effect on ROS; the SNs from ALS tissue increased the ROS.
That is, both tau and ALS SNs increased the ROS.
And now the right side of the figure, labeled QC-01-175. The same four conditions as on the left, but now they also add a drug, QC-01-175. The drug leads to the degradation of tau. All four bars of this set are low, similar to the control. In particular, the drug reversed the increase in ROS seen for two conditions at the left.
This is Figure 8b from the article.
|
Overall, the experiment shows that tau and ALS material can increase the production of reactive oxygen species, which are known to be harmful. A treatment that reduces tau reduces both of those effects. It is plausible that the effect of the ALS material on ROS was due to tau. It seems likely that, in ALS, tau is mis-localized to synapses, where it interferes with mitochondrial function.
Obviously, that experiment is not enough to make strong conclusions, but it fits with other evidence, from within this article as well as some background work. It is becoming clear that tau is important in the pathology of ALS. The phosphorylated form of tau affects mitochondria, in ALS as well as in other conditions.
And they have a drug that reduces tau. Clearly, they will be moving toward testing the drug to see if it affects the progression of ALS. That testing will start with animal models.
News stories:
* New Insights on ALS May Point To a Potentially Promising Treatment Strategy. (Neuroscience News (Mass General), November 10, 2021.)
* Tau Protein in Alzheimer's Linked to ALS Mitochondrial Dysfunction. (Marisa Wexler, ALS News Today, November 23, 2021.)
* Novel Target Identified for Treatment of ALS. (Ghazaleh Sadri-Vakili & Tiziana Petrozziello, Massachusetts General Hospital, December 27, 2021.) From two of the authors of the article.
The article: Targeting Tau Mitigates Mitochondrial Fragmentation and Oxidative Stress in Amyotrophic Lateral Sclerosis. (Tiziana Petrozziello et al, Molecular Neurobiology 59:683, January 2022.)
Recent post on ALS: Effect of exercise on developing ALS (July 27, 2021).
Recent post on tau: How many types of Alzheimer's disease are there? (June 15, 2021).
More tau: Selenium and neurodegenerative diseases? (June 25, 2022).
Posts about ROS include: How oocytes limit oxygen-induced aging (August 20, 2022).
More about brains is on my page Biotechnology in the News (BITN) -- Other topics under Brain. There is a list of related Musings posts.
February 16, 2022
Briefly noted... A mathematical analysis of egg shape
February 16, 2022
A recent article reports the ultimate egg equation, which can describe the shape of any type of egg. Key to getting the final, general equation was working out the equation for pyriform (pear-shaped) eggs.
* News stories; both link to the article...
- Research finally reveals ancient universal equation for the shape of an egg. (Sam Wood, University of Kent, August 31, 2021.)
- Why is an egg shaped like an egg? Turns out, there's some serious math behind it. (Mihai Andrei, ZME Science, September 16, 2021.) This news story includes the final equation.
* Direct link to the article: Egg and math: introducing a universal formula for egg shape. (Valeriy G Narushin et al, Annals of the New York Academy of Sciences 1505:169, December 2021.) Most of the mathematical development is in the Supplementary Material file, which seems to be freely available at the journal web site regardless of whether you have subscription access to the article itself.
* Also see: What is the proper shape for an egg? (September 18, 2017).
Briefly noted... A Swiss army knife for catalysis
February 15, 2022
A team of scientists has made a novel catalyst. It is, more or less, a random mixture of 14 metals, in a porous solid. That is, it contains multiple metals -- and sites with various combinations of metals. It actually works quite well in splitting water. And it could be useful in exploring novel reactions, since it provides a wide range of catalytic sites in a single material. How did they make it? Briefly, by making an alloy of the metals with excess aluminum, then dissolving away most of the Al. Their approach for making the stuff can probably be extended to other mixtures. It's all rather messy -- and intriguing.
* News story: Water splitting electrocatalyst made from unconventional alloy containing 14 elements. (Lucy Balshaw, Chemistry World, September 6, 2021.) Links to the article, which is open access.
* Direct link to the article (open access): Nanoporous ultra-high-entropy alloys containing fourteen elements for water splitting electrocatalysis. (Ze-Xing Cai et al, Chemical Science 12:11306, September 14, 2021.)
* Among posts on catalyst development...
- Added April 6, 2026.
Exploring the potential of aluminum(I) compounds as replacements for expensive metals (April 6, 2026).
- A tunable catalyst (July 25, 2022).
- A better catalytic sponge: degrading plastics, and more (August 11, 2020)
Is Kamo'oalewa a piece of the Moon?
February 14, 2022
Kamo'oalewa is a little thing (about 50 meters across) associated with Earth. It is perhaps best to say it is in orbit around the Sun, but it stays about ten million miles from Earth (about 40 times the distance of the Moon) -- at least for a while. It is called an asteroid, or a quasi-satellite (of Earth).
A recent article reports some measurements of its reflectance. Here are the main results...
The lines show reflectance (y-axis) vs wavelength (x-axis), for three known materials. The top line (blue) is for a Moon rock. The other two lines are for a couple of asteroids, considered fairly typical.
There are also some "loose" points, which are for Kamo'oalewa. They are close to the line for the Moon rock.
The spectra are normalized to 1 at 0.7 µm.
This is Figure 2 from the article.
|
There is more detail in the article, but that graph is the heart of the story. Kamo'oalewa looks like weathered silicate. In fact, it looks like lunar rock.
The authors discuss possible origins of Kamo'oalewa. There is little evidence to distinguish them, but one possibility is that it is a fragment of our famous big Moon, perhaps ejected during a collision. They are clear that this is a speculative proposal.
We noted at the top that Kamo'oalewa is rather close to Earth for a while. The following figure is a computer projection of its position, over several hundred years.
The y-axis is the distance from Earth, in AU (1 astronomical unit is the distance from Sun to Earth, about 90 million miles).
You can see that it is "close" to Earth, about ten million miles away, for a few hundred years. Before and after that period, it has a different orbit.
This is Figure 4c from the article.
|
News stories:
* Near-earth asteroid might be a lost fragment of the moon. (Science Daily (University of Arizona), November 11, 2021.)
* Piece of the moon? Asteroid might have lunar origin. (Paul Scott Anderson, EarthSky, November 18, 2021.) Several good pictures.
* Behind the Paper: Lunar-like silicate material forms the Earth quasi-satellite (469219) 2016 HO3 Kamo'oalewa. (Ben Sharkey et al, Nature Portfolio Astronomy Community, November 11, 2021.) From a group of authors of the article.
The article, which is open access: Lunar-like silicate material forms the Earth quasisatellite (469219) 2016 HO3 Kamo'oalewa. (Benjamin N L Sharkey et al, Communications Earth & Environment 2:231, November 11, 2021.)
For more about Earth's smaller moons...
* Briefly noted... 2. Earth's second moon (March 4, 2020). The current article mentions 2020 CD3, the mini-moon discussed in this earlier post, and includes two references to articles that have since been published about it (10 and 11).
* Briefly noted... 1. How many moons hath Earth? (September 5, 2018).
"Color vision" in mosquitoes?
February 12, 2022
Do mosquitoes have color vision? Are they attracted to the color of human skin? A new article provides some interesting tests.
The first figure shows the basics...
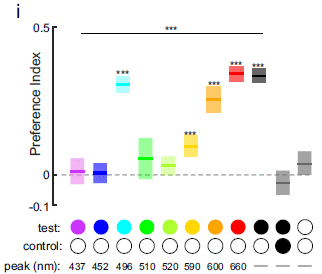
|
The plan is simple. The mosquitoes are offered two targets; the scientists observe which one they fly to. This leads to a "preference index", which is plotted on the y-axis.
Example... The first two tests, at the left, have choices between a white target and a purple or blue one. The preference index, from the testing, is zero.
Many of the results show zero preference. What is interesting is that some do not. The mosquitoes choose cyan, orange, red, and black over white.
The mosquitoes in this work are Aedes aegypti.
This is Figure 1i from the article.
|
We'll focus here on the orange/red part of that. Long wavelengths, and about the color of human skin.
Another test... In this case, the scientists knock out certain genes, and test if that affects the mosquitoes' preference for the reddish colors.
The first result is for the wild type (wt) mosquitoes. They show a preference for red, as expected from the first test.
The important result is that two mutant strains do not show that preference.
One of them is at the far right. It is a double mutant, in which two types of visual pigments (opsins) have been knocked out. These two opsins are thought to respond to long-wavelength light. Knocking them both out eliminates the response to long-wavelength light. (As controls, the two strains each lacking one of those visual pigments retain the response.)
|
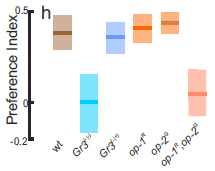
|
The other mutant strain that does not show the red-preference is labeled Gr3(-/-). (The heterozygote (+/-), just to the right, retains the preference.) This implicates the Gr3 gene in the color preference response. What does Gr3 do? It is part of the system for detecting CO2.
This is Figure 2h from the article.
|
The general picture that emerges from the work...
- Mosquitoes have some color vision.
- Mosquitoes are attracted to the color of human skin.
- Mosquitoes respond to the color of human skin only when it is accompanied by the odor of carbon dioxide.
It is that last point that is of particular interest.
Those conclusions are for Aedes aegypti. Two other types of mosquitoes studied briefly in this work also showed that visual responses were affected by CO2, but the details varied.
That mosquitoes smell CO2 had been established previously. (Humans do not smell it.) The finding here is a connection between smelling something and seeing something.
There has been previous work on color vision in mosquitoes, with inconsistent results. An interesting point is that the basic vision testing in the current article was done while controlling the CO2 level.
News story: New Mosquito Vision Discovery Could Help You Hide From These Disease-Carrying Bloodsuckers. (SciTechDaily (University of Washington), February 4, 2022.)
The article, which is open access: The olfactory gating of visual preferences to human skin and visible spectra in mosquitoes. (Diego Alonso San Alberto et al, Nature Communications 13:555, February 4, 2022.)
A post on mosquito olfaction, which notes CO2: Why don't black African mosquitoes bite humans? (December 19, 2014).
This post is listed on my page Internet resources: Biology - Miscellaneous under Medicine: color vision and color blindness. It includes links to several Musing posts. Added June 4, 2025.
Posts on the general nature of mosquitoes are included in the listing at a section of my page Biotechnology in the News (BITN) -- Other topics on Malaria.
February 9, 2022
Extraction of rare-earth metals: a bacterial assist
February 8, 2022
A recent article reports progress toward an improved method for extracting rare earth elements (REE) from minerals. The progress is modest, but the approach is interesting and more general.
The proposed process makes use of bacteria of the genus Gluconobacter. These bacteria have the special property of oxidizing glucose to gluconic acid, thus acidifying the growth medium. The resulting acidity helps dissolve the mineral sources of the REE, thus bringing the REE ions into solution. It could be an environmentally friendly process, but it is not well understood, and there has been little development. (The full story of how the culture fluid solubilizes the REE is probably more complex than just the gluconic acid.)
The general approach of the new work is to look for mutant bacterial strains that make more acid, and then test them to see if they are better at dissolving REE minerals. The mutant hunt starts by making a set of gene knockouts: a collection of strains in which each member has been mutated in one of 2700 genes of the bacteria. (Only non-essential genes can be tested using this method.) Some effort is required to make such a "library", but once it is made, it can be used for various purposes.
The scientists did two screens for acid production. One looked for mutants that made the final pH unusually low. The other looked for mutants with a high initial rate of acid production. The following figure shows some results from the second screen.
The graph shows the initial rate of acid production for some of the strains. The green bars are for mutant strains with the highest such rates. A few of the worst strains are shown at the left in blue.
A few strains have substantially higher rates of acid production. They can be studied further.
This is Figure 2E from the article.
|
The following figure illustrates the possible benefit of such strains. It also illustrates the challenge.
In this lab-level test, several bacterial strains were tested to see how they solubilized an REE mineral-like material. The graph shows the amount of REE solubilized (y-axis) vs the culture pH (x-axis).
It is clear that there is a good trend. Lower pH does indeed lead to better solubilization of the REE.
Biolixiviant? It refers to the culture fluid from the bacteria. The cells have been removed.
This is Figure 4E from the article.
|
Closer examination reveals some limitations. The original (wild type) strain is at pH 2.3. Some of the new mutant strains do indeed give better extraction of the REE, but the improvement is relatively small. And the high variability leads to uncertainty about the significance.
Taken at face value, the best mutant strain increases the REE extraction by 18%. Is that enough to be useful? That is not clear.
However, questioning the statistical significance or importance of the current results may be missing the point. This is step 1. The scientists have identified some strains, many with mutations in known genes, that lead to lower pH and better REE extraction. Further work can explore using combinations of those mutations. And the knowledge gained from these strains may aid in the rational design of an improved process.
News story: Bacteria could extract elements for modern tech sustainably. (Krishna Ramanujan, Cornell, November 18, 2021.)
The article, which is open access: Generation of a Gluconobacter oxydans knockout collection for improved extraction of rare earth elements. (Alexa M Schmitz et al, Nature Communications 12:6693, November 18, 2021.)
A follow-up post on this work: Using bacteria to leach valuable metals, including rare earths, from rocks -- and to capture atmospheric carbon (September 4, 2025). Added September 4, 2025.
Posts about rare earth elements include:
* Briefly noted... A practical way to make tetrataenite -- and reduce the demand for rare earth elements (January 25, 2023).
* Briefly noted... A terbium detector (January 19, 2022).
* Penidiella and dysprosium (September 11, 2015). Links to more.
This post is listed on my page Introductory Chemistry Internet resources in the section Lanthanoids and actinoids. That includes the REE.
Communication in diatoms
February 6, 2022
A recent article suggests that diatoms in the ocean send signals around the colony using flashes of red light.
Let's look at a couple of pieces of the evidence. Caution, the story in the article is not very complete. But it is a fun story.
The figure shows how diatoms change shape as they are irradiated with blue light.
The y-axis shows R, a measure of the shape of the cell. (It is the ratio of the lengths of two axes of the cells.)
The blue curve shows the results for live cells. The cell shape oscillates. As a control, dead cells don't respond (black curve).
The red line is a sine wave that is a good fit to the results.
This is Figure 1A from the article.
|
That is, shining continuous blue light on the diatoms causes the cell shape to oscillate. The wavelength of the oscillation is about 100 seconds.
The next figure shows what happened if the scientists gave pulsed red light...
Again, the y-axis shows the cell shape (R). Blue curve; left-hand y-axis.
For the first half of the test, there is no light. The cell shape varies, seemingly randomly.
At about 1400 seconds, the red light is started, in pulses. Red curve; right-hand y-axis, though the numbers are not important for now.
The blue and red curves follow each other rather closely. That is, the cell shape follows the pulsing of the red light. It's certainly not perfect, but the difference between the two halves of the figure is clear.
The period is about 100 s, as in the first experiment. In fact, the test works best if that period is used for the light pulses. (100 s period = 0.01 Hz frequency.)
This is Figure 3A from the article.
|
A few details...
- Blue light penetrates water better than red light.
- It is known that the diatoms have a receptor for red light. Why has been a mystery.
- The chlorophyll in the diatoms can fluoresce, giving red light.
Putting the pieces together, the authors come up with a model... Blue light, which can penetrate the water, leads to red fluorescence from the diatom cells. The diatoms sense the red fluorescence from their neighbors, and use it to change their cell shape, in synchrony.
How do we get from continuous blue light (in nature, sunlight) to pulsed red light (from the diatoms)? The authors argue that this is due to the geometry of the colony, and how the cells are aligned.
What's novel is that it is commonly assumed that single-cell organisms communicate largely with chemical signals. Here, it is proposed that they send and receive light signals.
Why do the diatoms do this? That must be left for further work. One possibility is that the oscillating alignment improves the efficiency of use of light for photosynthesis.
If you aren't convinced by all that, that's fine. The article offers preliminary results about an unfamiliar system. Sometimes, that is the fun part of science, opening up something new and unusual.
News stories:
* Evidence found of diatoms communicating with each other using natural fluorescence. (Bob Yirka, Phys.org, December 17, 2021.)
* Diatom Dialogue. (Fayth Tan, oceanbites, January 27, 2022.) (The statement referring to "sensory cells" should say "sensory molecules".)
* Q&A: Fluorescence Lets Diatoms Communicate, Coordinate Behavior -- The Scientist spoke with physicist and microbial ecologist Idan Tuval, whose recent paper challenges the assumption that these single-celled organisms only communicate via chemical signals. (Dan Robitzski, The Scientist, December 16, 2021. Now archived..) Interview with one of the authors.
The article, which is open access: Pelagic diatoms communicate through synchronized beacon natural fluorescence signaling. (Joan S Font-Muñoz et al, Science Advances 7:eabj5230, December 15, 2021.)
More about diatoms:
* Item #1 at Briefly noted... 1. Underwater landslides (August 15, 2018).
* Fertilizing the ocean may lead to reducing atmospheric CO2 (August 24, 2012).
* Titanium biology (September 29, 2008).
Also see: Physiological effects of blue light (September 10, 2022).
FGF1 and insulin?
February 5, 2022
Insulin is a hormone that helps to control the level of glucose in the blood. Recent work has uncovered a second factor that seems to play about the same role.
The factor is called fibroblast growth factor 1 (FGF1). Work in mice has shown that it has anti-diabetic effects, but how it works has been unknown. A new article explores that question.
A specific role of insulin is to inhibit lipolysis, the breakdown of fats. The first figure looks at the role of FGF1 in this process...
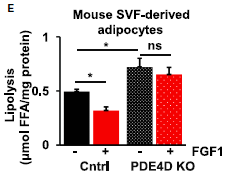
|
The bar heights show the degree of lipolysis, under different conditions. It is shown as the amount of FFA (free fatty acids) released. This is a lab test, using adipocytes (fat cells) from two types of mice.
The first two bars (left) are for adipocytes from ordinary (control) mice. The right-hand bar of the pair (red) shows what happened when FGF1 was added. There was a significant reduction in FFA released. FGF1 inhibits lipolysis.
|
The next two bars (right) are for adipocytes from a mutant mouse strain, labeled PDE4D KO. KO means knockout; the gene in question has been inactivated. The gene is a PDE = phosphodiesterase; it is number 4D of a large set of such enzymes.
Knocking out PDE4D led to a substantial increase in lipolysis. And adding FGF1 (red bar) had little effect.
This is Figure 4E from the article.
|
That is, the results of that experiment show that FGF1 inhibits lipolysis, and that it acts via enzyme PDE4D.
That is just one piece of the story. The following figure summarizes what the scientists think FGF1 does in this context, and compares it to what insulin does.
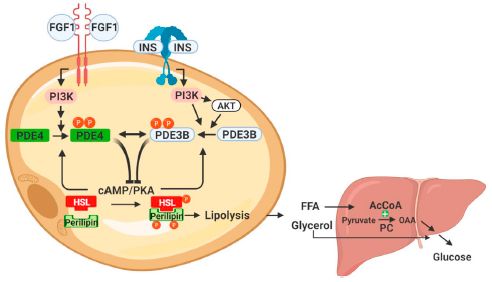
In this cartoon figure, the big object at the left is an adipocyte; the object to the right is a liver.
Look at where they are near each other. Lipolysis in the fat cell leads to the production of FFA, which enter the liver (and lead to more glucose). That lipolysis is the process of interest.
Backing up... Lipolysis requires cAMP/PKA, as shown just to the left. (We'll skip identifying what all those terms mean.) But just above the label "cAMP/PKA" two lines come in as inhibitors. The small perpendicular end on a line indicates a block.
At the top are FGF1 and INS (insulin). Those two factors lead down to those two blocks at cAMP/PKA. That is, both FGF1 and INS do about the same thing: inhibit lipolysis.
The difference? FGF1 acts through PDE4D, as shown in the first figure here. Insulin acts through another member of the PDE family, PDE3B.
This is Figure 6 from the article.
|
That's the big idea from the article: FGF1 and insulin act in parallel to affect lipolysis.
Insulin is therapeutically useful. Should FGF1 be useful, too? That question is now open.
News stories:
* Molecule Produced in Fat Tissue Could Unlock New Approaches to Diabetes Therapy. (GEN, January 5, 2022.)
* New Route Discovered for Regulating Blood Sugar Levels - Independent of Insulin. (SciTechDaily (Salk Institute), January 10, 2022.) A cute cartoon at the top captures the key point. Also includes an interesting COVID-appropriate picture of part of the team of scientists.
The article: FGF1 and insulin control lipolysis by convergent pathways. (Gencer Sancar et al, Cell Metabolism 34:171, January 4, 2022.)
Previous post about insulin: A gastric auto-injector, which gives shots in the stomach lining (January 24, 2022).
More on diabetes is on my page Biotechnology in the News (BITN) -- Other topics under Diabetes. That includes a list of related Musings posts.
February 2, 2022
Briefly noted... How and why worms spit
February 2, 2022
Have you ever watched a worm spit? Do you know why a worm would spit? Why shining light on a worm might lead to it spitting, even though it lacks light receptors? If you are inclined to not take this seriously, we should note that the work has revealed some new ideas about how a muscle cell works. Amazing what one can do with C elegans.
* News story: News Brief: Mapping the cellular circuits behind spitting -- Roundworms change the flow of material in and out of their mouths in response to bright light, revealing a new way for neurons to control muscle cells. (Raleigh McElvery, MIT, July 23, 2021.) Includes two videos, one an action shot and one a narrated overview of the work. Links to the article, which is open access.
* A recent post on C elegans as a model organism: Lesson from a worm: An endogenous anti-opioid system? (January 5, 2020). Links to more.
Was hydrogen the energy source for the origin of life?
February 1, 2022
The following figure, from a recent article, offers an important insight into the nature of life...
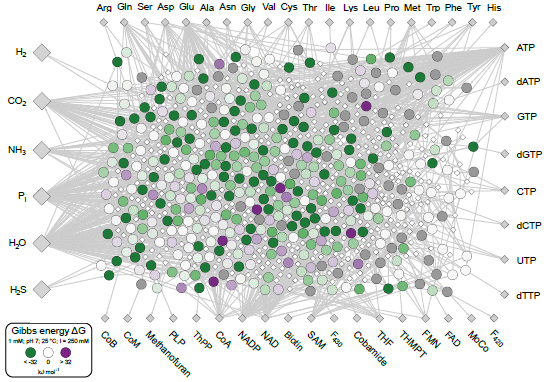
That's a complicated figure!
Let's see if we can extract one key idea from it, and avoid a detailed explanation.
Key observation: There are more green circles than purple circles.
Each circle represents one biochemical reaction. The collection of circles (about 400 of them) represents the core metabolism of biology. They are reactions that are almost universal, and thus we might infer were probably present in very early cells.
The color indicates whether the reaction is favorable. Green-circle reactions are favorable. Purple-circle reactions are unfavorable. The color intensity shows this quantitatively. The color intensities in the key are for ΔG of +/- 32 kJ/mole. We might amend our first observation, that there are more green circles, to say that there are more dark green circles than dark purple circles. Far more. (The ΔG values are for specific conditions, thought relevant here, at least as a baseline case. Gray circles: no value is available.)
The general layout of the figure is that the building blocks are shown along the left side, and various classes of bio-molecules along the other three sides.
This is Figure 1 from the article.
|
The conclusion? Most biochemical reactions are energetically favorable. That might suggest that the origin of life was an energetically favorable event (not requiring an external energy source, such as light).
Is there a catch? You bet. The conclusion depends on the starting conditions, including what chemicals are used, as well as environmental conditions such as temperature and pH.
So, what is the key condition? The presence of hydrogen gas, H2. We all know that hydrogen is a good fuel. If life formed using H2, it could be that there was no need for any further energy source. That is the main message from the article.
Is it reasonable that life began with H2 around? Deep-sea hydrothermal vents have long been considered as reasonable sites for life to begin. The thermal energy of such sites is an appareling feature. H2 is known to be abundant at some such sites. So, yes, it is reasonable that life could have started using H2, both as a building block and as a fuel.
The analysis in the article can only be taken as a clue. There is no proof of what happened. And the analysis is incomplete; it does not consider polymer formation, and it is based on a cell that is some distance away from the first life. But the conclusion is interesting and reasonable; the possibility that the original energy source for life was hydrogen gas belongs on the table.
The article is rather difficult reading. Both news stories listed below are from the authors and the main institution; they are good places to start.
News stories:
* Life arose on hydrogen energy. (EurekAlert! (Heinrich-Heine University Duesseldorf), December 13, 2021.)
* Likely energy source behind first life on Earth found 'hiding in plain sight'. (Jessica Wimmer & William Martin, Science & research news, Frontiers, January 19, 2022.) From two of the authors, in a blog from the journal publisher.
The article, which is open access: Energy at Origins: Favorable Thermodynamics of Biosynthetic Reactions in the Last Universal Common Ancestor (LUCA). (Jessica L E Wimmer et al, Frontiers in Microbiology 12:793664, December 13, 2021.)
A post about hydrogen-based life near a thermal vent: The hydrogen economy -- in the mid-Atlantic (August 30, 2011).
More about the possibility of hydrogen as a food: Is there food on Enceladus? (May 21, 2017).
More hydrogen: Making hydrogen visible (July 2, 2022).
A recent post about origin-of-life chemistry: Miller-Urey revisited: the role of the glass container (January 22, 2022). Last week. Links to more.
more... Making peptides in space: a new pathway (April 15, 2022).
Triclosan: an explanation for its gut toxicity
January 30, 2022
The antimicrobial agent triclosan has long been the subject of controversy. It is a common disinfectant, and has been incorporated into many consumer products. It is now partially restricted in the US, but questions remain. Triclosan and its problems were introduced in an earlier post [link at the end].
The current concern is the association of triclosan with gut inflammation (colitis). There is a lot of evidence of a problem, sometimes serious, but how it occurs has not been clear. A new article offers some insight.
The following figure lays the groundwork for the new story...
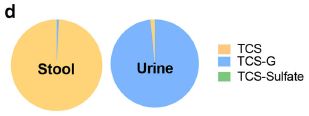
|
The pie charts show the forms of triclosan excreted by humans.
The quick observation is that triclosan is excreted in different forms in stool (feces) and urine.
The orange form, the major form in feces is triclosan itself (TCS). The blue form, the major form in urine, is TCS attached to glucuronic acid (TCS-G). (The key shows a third form, which is minor.)
This is Figure 1d from the article.
|
What's going on? Attachment to glucuronic acid is a common way to mark a substance for excretion. In fact, the formation of TCS-G was well known; it was assumed that was the major pathway for getting rid of TCS. The results above suggest the story is more complicated.
Note that the figure above does not give the relative amounts excreted by the two pathways. The important point is that the form excreted in feces is different from what was expected.
The following figure provides some information about what happens...
The left side shows the chemical structures of TCS-G (top) and TCS (bottom).
The context is that gut bacteria break down the TCS-G to make free TCS.
The right side provides some evidence for that conversion. The bars show the percent of free TCS in three samples. The first is for a control sample of TCS-G; it has a very low amount of free TCS. The next two samples have been treated with a mixture of fecal bacteria from mouse (middle bar) or human (right). In both cases, the bacteria led to a substantial production of free TCS.
This is Figure 2a from the article.
|
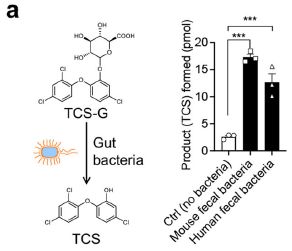
|
Other tests showed that antibiotics led to lower levels of free TCS, again implicating the gut bacteria.
The enzyme activity needed to convert TCS-G to free TCS is a beta-glucuronidase (GUS). It is known that the gut microbiota contains many types of such enzymes, with various specificities. The scientists here showed that some of them indeed convert TCS-G to TCS. An inhibitor of those enzymes reduced the conversion, and in vivo reduced colitis in mice fed TCS.
Overall, the article provides considerable evidence that TCS-G is converted to free TCS by β-glucuronidase enzymes from gut bacteria, in both mice and humans. Further, it appears that free TCS is responsible for the gut inflammation associated with triclosan.
News stories:
* Animal Study Demonstrates How Triclosan Can Trigger Gut Inflammation. (Sci-News.com, January 11, 2022.)
* Gut Bacteria Enzymes Link Antimicrobial Triclosan with Gut Inflammation, IBD. (GEN, January 10, 2022.)
The article, which is open access: Microbial enzymes induce colitis by reactivating triclosan in the mouse gastrointestinal tract. (Jianan Zhang et al, Nature Communications 13:136, January 10, 2022.)
Background post: Does Triclosan in antibacterial soaps promote infection? (May 19, 2014).
Recent posts about gut microbiomes:
* Could a better gut microbiome improve memory? (December 6, 2021).
* Treating malnutrition by improving the gut microbiome (May 29, 2021).
My page for Biotechnology in the News (BITN) -- Other topics includes a section on Antibiotics. It includes a list of Musings posts on the topic, including disinfectants.
The effect of Starlink satellites on astronomical observations
January 29, 2022
The following figure features the Andromeda galaxy, as seen with a telescope at Mount Palomar. It also features a human-made object that got in the way.
Note the near-vertical streak about 1/3 of the way from the right edge. A communications satellite passed by.
The image is from a sky survey project called Zwicky Transient Facility. It uses a telescope at Palomar, operated by Caltech.
This is from the Caltech news release listed below.
|
The following figure shows how often that happens...
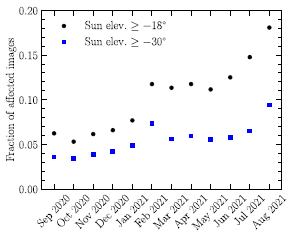
|
The graph shows the fraction of images with such streaks over a recent one-year period. Results are shown for two sun-elevations.
The big picture is that the frequency of affected images has risen, and is now nearly 20%.
Further, the problem is worse at twilight, with the sun just over the horizon (upper data set). Twilight is an important time for the project; it is when asteroids near the Earth are best observed.
This is Figure 2 from the article.
|
Satellites are getting smaller and cheaper -- and more numerous. The current satellites are part of the Starlink system, from SpaceX. The first went up in 2019. At the end of the period shown above, about 1600 were in low-Earth orbit. The number of such satellites approximately doubled during the time covered above (see Figure 1 of the article). Plans have been announced for many thousands more; some people estimate that there may be a hundred thousand of them within a decade.
Astronomers have been concerned about the possible effects of the satellites on their work. The results above provide some evidence about the current state.
SpaceX has offered changes to reduce the problem. One change was implemented, but then withdrawn because it affected the satellites adversely. Another is being tested; results reported in the article show that "visors" make a small but significant improvement. Software can process out the satellite streaks; after all, the satellites have known orbits. But the problem will become worse as the number of satellites increases. Further, other telescope systems may be affected more.
It is a clash of technologies. And it is not yet resolved.
News stories:
* Starlink Satellites Don't Impact Science (Yet). (Monica Young, Sky & Telescope, January 25, 2022.)
* Palomar Survey Instrument Analyzes Impact of Starlink Satellites -- A study of archival images from Zwicky Transient Facility shows an increase in satellite streaks. (Whitney Clavin, Caltech, January 17, 2022.)
The article, which is open access: Impact of the SpaceX Starlink Satellites on the Zwicky Transient Facility Survey Observations. (Przemek Mróz et al, Astrophysical Journal Letters 924:L30, January 14, 2022.)
For more about small satellites, including the Starlink system, see the news feature noted as #2 at Briefly noted... 2. CubeSats (November 7, 2018). Starlink satellites are not the tiny ones most often associated with the term CubeSat, but they are part of the trend toward smaller satellites.
January 26, 2022
Briefly noted... Enzymes for degrading plastics may correlate with presence of plastics
January 26, 2022
A recent article reports a vast global survey of environmental DNA looking for genetic signs of plastic-degrading enzymes. The scientists find a correlation with the presence of plastics in the environment. Is this a sign that plastics are being increasingly degraded in Nature? It is a complex and intriguing article. Regardless of the claims, it should lay the groundwork for future studies. In particular, it may point to the availability of new enzymes.
* News story: More microbes that can degrade plastics in places with heavy plastic pollution. (Science Daily (Chalmers University of Technology), December 14, 2021.) Links to the article, which is open access.
* Among posts on degradation of plastics... Good enzymatic degradation of polyesters, by manufacturing the plastic with the enzymes in it (May 4, 2021).
Briefly noted... How big were medieval war horses?
January 25, 2022
Probably not as big as the common view, according to a new article. It is an interesting archeological exploration, published in the International Journal of Osteoarchaeology. (Horses are measured in "hands"; 1 hand = 4 inches. Figure 2 of the article is labeled in both meters and hh = hands high; 1.4 m is about 14 hh. Horse height is measured to the withers, or shoulder; it is the high point of the back, and does not include the head.)
* News stories. Both link to the article, which is open access.
- Medieval warhorses were surprisingly small in stature. (Science Daily (University of Exeter), January 10, 2022.)
- Pony-sized horses carried medieval knights into battle, research shows. (Horsetalk, January 13, 2022. Now archived.)
* More horse history...
- Added November 5, 2025.
Why horses are rideable: GSDMC (November 5, 2025).
- How horses learned to walk (September 21, 2016). Links to more.
A gastric auto-injector, which gives shots in the stomach lining
January 24, 2022
Don't like folks sticking a needle in your arm? Perhaps you could swallow the needle, and take the shot in your stomach.
That's not exactly what happens here -- but it is close. It does give you an idea of a new approach to injections, as recently reported.
Here is what you swallow...

|
It is a SOMA, specifically an L-SOMA = liquid-injecting self-orienting millimeter-scale applicator.
It is 1.5 centimeters high -- near the limit of what is allowed. (Yes, there are recognized limits.)
The scale bar? It is not identified, but I suspect it is 0.5 cm.
This is Figure 1e from the article.
|
The following figure is a cartoon diagram of what the L-SOMA does in your stomach. The picture of the L-SOMA here in based on a CAD drawing, which you can see in a larger form in Figure 1c of the article.
First, the SOMA orients itself against the stomach lining (the wavy structure at the bottom).
Second, part of the red plug near the top dissolves away. You can see the dissolved red stuff loose at the top. That leads to the needle in the SOMA inserting itself into the stomach lining. You can see the needle sticking into the stomach lining.
Third, the dose is delivered. You can see the blue dose-stuff in the stomach lining. Where did it come from? It is stored in a chamber very near the bottom. You can see some blue there is the first frame. (You can see it better in the larger figure in the article.)
Finally, the second piece of red-stuff is dissolved away, and the needle retracts back into the SOMA. It can now be excreted.
This is Figure 1a from the article.
|
Here are some results...
The test here is giving insulin to pigs.
Part a (upper left) shows the insulin levels obtained. Don't worry about sorting out the lines; the differences are not important for now. But for the record... The blue lines are for individual pigs; the black lines show the averages. The two symbols (dashed vs solid) are for two variations of how anesthesia was given. We'll come back to that.
In part b (upper right), the blue curves show the resulting blood glucose levels. The upper lines (green) are for negative controls (no insulin). The black lines show the averages for each set. You can see that the insulin reduced the glucose levels.
Parts e and f (lower) show insulin and glucose levels following ordinary insulin injections.
This is part of Figure 3 from the article.
|
It worked. A high percentage of the dose was delivered with good kinetics. Blood sugar levels decreased.
There is no claim that the results with the new device are better. In fact, they are less consistent than the traditional injections (and there is an occasional failure, not seen in the graphs above). But the ability to give protein drugs orally is appealing. Protein drugs are usually injected; taking them orally would lead to digestion in the stomach. Taking a "pill" or capsule is easier; the current L-SOMA has the potential of allowing protein drugs to be given orally, yet avoiding stomach digestion.
In the pig work, the scientists placed the capsule in the stomach, using an endoscope; they also gave the pigs an anesthetic during this procedure. They briefly note that they have used the device with dogs, with the SOMA capsule given orally without an anesthetic. Presumably, the anesthetic is not an important part of the procedure here.
They tested a range of drugs, from small metabolites to large antibodies. The L-SOMA worked for all molecules tested. Pills are better than injections, from the point of view of both the patient and health-care staff who administer them. And for delivering large quantities of large molecules such as antibodies, infusion may be needed, making it even more complex. Given the potential advantages, the scientists want to move toward testing the new L-SOMA in humans.
News stories:
* Self-injecting pill allows for oral administration of monoclonal antibodies and other injected drugs -- In pre-clinical testing, traditionally injected drugs such as insulin, epinephrine, GLP-1 receptor agonists and monoclonal antibodies were orally delivered with comparable results. (EurekAlert! (Brigham and Women's Hospital), August 30, 2021.)
* Drug delivery capsule could replace injections for protein drugs -- The new pill can inject large quantities of monoclonal antibodies and other drugs into the lining of the stomach after being swallowed. (Anne Trafton, MIT, August 30, 2021.)
The article: Oral delivery of systemic monoclonal antibodies, peptides and small molecules using gastric auto-injectors. (Alex Abramson et al, Nature Biotechnology 40:103, January 2022.)
More insulin... FGF1 and insulin? (February 5, 2022).
More about vaccine technologies: Self-boosting vaccines? (August 26, 2022).
This post is listed on my page Biotechnology in the News (BITN) -- Other topics in the section Vaccines (general). The current post may not be about vaccines per se, but it is related. And the method could be used for vaccines, too. The section has a list of Musings posts on various vaccine issues, including alternative delivery modes.
Miller-Urey revisited: the role of the glass container
January 22, 2022
In 1953 Stanley Miller, working with Harold Urey, published an experiment in which they took a mixture of simple chemicals, provided a spark, and found more complex chemicals, including amino acids. They had made biotic chemicals from abiotic chemicals. It was the first definitive experiment to show this; the article became "Exhibit 1" in the new field of origin-of-life chemistry.
The details of Miller-Urey have been argued about over the years. Yes, it is interesting, but how relevant were the conditions?
Interestingly, one aspect of Miller-Urey has not been discussed much, They used a glass container. Borosilicate (Pyrex) glass. Does that matter? A recent article addresses that question.
Here is the answer...
The two tubes at the left are from a test using glass. The middle tubes are from the same type of test but run in a Teflon container. (The right-hand set is from a test using a Teflon container, but with bits of glass added. We won't pay much attention to the results in that system.)
You can see that the test run in the glass container gave more brown/yellow stuff in solution, and more stuff at the bottom.
This is Figure S2 from the Supplementary Information with the article.
|
That's a clue, but the scientists went on to do more complete analysis. The following table summarizes what they found...
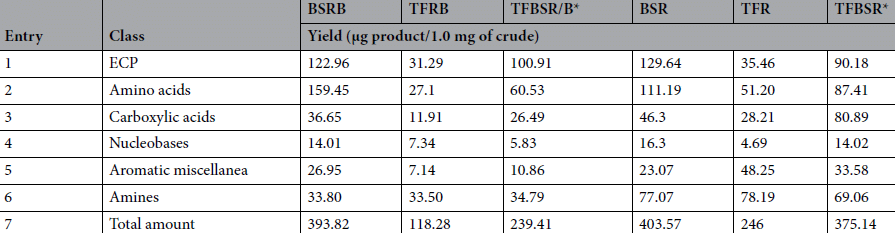
The table shows the amount of chemicals of various types made under various test conditions.
The test conditions are based on the three conditions shown above, but with one more variable: pH. The three columns at the right are for unbuffered solutions, which started at about pH 11. The three columns at the left are for the same conditions, but now with a buffer, at pH 8.7. Higher pH should increase the amount of silicate that dissolved.
For a start, look at row 7 (bottom) of the table, for the total amount of products, in micrograms products per milligram of the starting chemicals.
For both sides (both pH's), there was more product in glass than in Teflon. The test with Teflon-with-glass gave intermediate results. The results for buffered and unbuffered conditions were similar, though you can find some interesting differences in detail.
The first six rows are for types of products. (ECP = elemental prebiotic chemical precursors.) The results for amino acids are perhaps the most dramatic.
About a third of the chemicals that were characterized were made only with glass.
BSR = borosilicate reactor. TFR = Teflon reactor; BS at the end means added borosilicate. For each reactor type, a B at the end indicates buffer.
This is Table 1 from the article.
|
The big story here is that the nature of the container matters. The Miller-Urey experiment gave as good a result as it did partly because they used a glass container.
That's not a big surprise. Much work has shown how useful silicate surfaces are in catalyzing reactions. Many people have speculated that mineral surfaces were likely important in the reactions that led to life. It may be beyond us to ever establish what happened 3+ billion years ago, but the current work does add an interesting piece to a famous origin-of-life experiment.
News stories:
* Glassware found to promote reactions in Miller-Urey 'primordial soup' experiment. (Katrina Krämer, Chemistry World, November 8, 2021.)
* Scientists recreated classic origin-of-life experiment and made a new discovery -- 1952 Miller-Urey experiment showed organic molecules forming from inorganic precursors. (Jennifer Ouellette, Ars Technica, October 28, 2021.)
The article, which is open access: The role of borosilicate glass in Miller-Urey experiment. (Joaquin Criado-Reyes et al, Scientific Reports 11:21009, October 25, 2021.)
Posts about origin-of-life chemistry that mention the Miller-Urey experiment:
* Modeling the role of hydrogen cyanide in the pre-biotic formation of life's chemicals (September 8, 2019).
* The origin of life's chemicals: making an almost-Krebs-cycle in the lab (June 10, 2019).
More origin-of-life chemistry:
* A special role for cyanide in the chemistry of early life? (April 25, 2022).
* Was hydrogen the energy source for the origin of life? (February 1, 2022).
More about silicates: Silicates and gene expression -- a new way to induce making cartilage? (June 21, 2022).
More Teflon: Purifying water using fluorinated nanopores (August 16, 2022).
January 19, 2022
Briefly noted... A terbium detector
January 19, 2022
This may be mainly of interest to those who use terbium (element 65; Tb) in their daily lives. That would include most anyone who uses a phone. Tb is a rare-earth, or lanthanoid, element, and a relatively rare one at that. The new work builds a sensitive Tb detector with high specificity. It is based on the recently discovered natural protein called lanmodulin. The scientists modified the protein, to enhance its fluorescence when Tb is bound. The understanding gained during the development may help the scientists make detectors for other rare elements.
* News stories; both link to the article:
- Luminescent Sensor Identifies Valuable Rare Earth Element Terbium in Unexpected Locations. (SciTechDaily (NSF), December 13, 2021.) Brief; mainly on the use of the device for finding Tb in environmental sources.
- New sensor detects valuable rare earth element terbium from non-traditional sources. (Gail McCormick, Penn State, August 25, 2021.)
* More about lanmodulin: A protein that can assist with handling actinium (January 9, 2022).
* Posts about rare earth elements include...
- Extraction of rare-earth metals: a bacterial assist (February 8, 2022).
- Penidiella and dysprosium (September 11, 2015). Links to more.
* This post is listed on my page Introductory Chemistry Internet resources in the section Lanthanoids and actinoids.
What caused the extinction of the mammoths -- II
January 18, 2022
This is the second of two consecutive posts on the topic. The first is immediately below.
The article here, like the first, does statistical analysis of large data sets. The following figure summarizes the findings for one Arctic region.
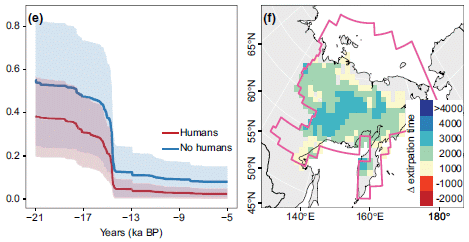
This graph is for Beringia, the region from northwest Siberia to Alaska.
Part e (left) shows estimates of the size of the mammoth population (in millions; y-axis) over time (x-axis), using models with and without humans (red and blue, respectively). (ka BP = kilo-years before present.)
Big picture: By these analyses, humans substantially reduced the population of mammoths. Numerically, the greatest effect was in the early days.
Part f (right) shows the effect on a local level. The modeling yields estimates of when local extinction (called 'extirpation') would occur, with and without humans. The difference between those two numbers is shown here. Positive values (blue/green/yellow) mean the population would have lasted longer without humans. Negative values (red) mean that the population would have lasted longer with humans; there are no red pixels on this map (I think).
This is part of Figure 5 from the article. The full figure also includes such analysis for two other regions of the North; the general picture is the same, though there are interesting geographical variations.
|
Another big analysis. And this one suggests a significant role for humans in the population decline of the mammoths. In particular, it shows a significant effect of humans on the mammoths rather early (by the time of the LGM (Last Glacial Maximum). The authors argue that the early population decline had continuing consequences. That is, the mammoth populations were smaller at all times, thus making the ultimate extinction from very small population sizes occur earlier. On the other hand, we might note that the biggest population decline seen in the figure, at about 14 ka ago, is not due to humans.
What is the analysis here? It uses generally available data. The main advance, the authors argue, is that they build models around ecological processes, rather than simply looking for statistical correlations. That leads them to make a causal connection between the early population decline and the later extinction.
Some final comments about the pair of related posts...
The headlines about them emphasize that they reach opposite conclusions about the role of humans in the extinction of the mammoths.
Both articles are hugely complex. Both depend on complex statistical analysis of large complex data sets. One article introduces an important new data set, the order introduces a complex approach to the analysis
As you look more closely, the two articles emphasize different parts of the story of the mammoths. The first article says that humans were not a major player in the last few thousand years of the mammoths. The second article says that humans were an important factor earlier, and that the early effect had later consequences. It is quite possible that both are right. But it is also possible that both analyses are incomplete; most work in this field is incomplete.
These are both interesting articles, which make significant contributions. But try to go beyond the headlines, and see what they actually did. A simple headline gets attention, but is not always the full story of what an article is about. Musings posts are more than headlines, but they are brief presentations of a portion of what was done.
News stories:
* Study: Humans Played Significant Role in Extinction of Woolly Mammoths. (Sci-News.com, November 11, 2021.)
* Mammuthus Primigenius: Human Activities Speed Up Extinction of Ancient Woolly Mammoths, Study Says. (Ron Jefferson, Science Times, November 13, 2021.) This news story makes some attempt to connect the articles of our two posts here. Unfortunately, it is not well-written, so be patient reading it.
The article: Process-explicit models reveal pathway to extinction for woolly mammoth using pattern-oriented validation. (Damien A Fordham et al, Ecology Letters 25:125, January 2022.) The article is currently "free to read" at the given link; you cannot download or print it.
The first post of this set: What caused the extinction of the mammoths -- I (January 16, 2022). Preceding post, immediately below. It links to other related posts. The article of that post is referenced in the current article, as Wang et al (2021).
What caused the extinction of the mammoths -- I
January 16, 2022
There are two related posts on this topic. The second is immediately above. Each is presented separately, with a brief overview. At the end of the second post, there is a brief comparison. I urge readers to reserve judgment until you have seen both posts.
Megafauna, such as the woolly mammoth, went extinct a few thousand years ago. Why? Many possible factors have been suggested, but we don't know. Of particular interest is the possible role of humans in the extinction of the megafauna? Did humans kill them off?
Here we address the first of two recent articles addressing the issue.
The heart of this new work was collection of a large number of samples of eDNA -- environmental DNA -- from samples throughout the Arctic. Analysis of this eDNA yielded evidence for what plants and animals were present at various times over the last 50,000 years. It is another example of metagenomics. The scientists then analyzed that biodiversity data along with climate records.
Here is an example...
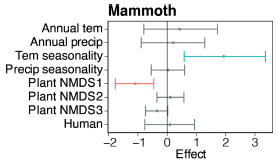
|
The figure shows how various factors, listed at the left, appeared to affect the population of the mammoths.
tem = temperature; precip = precipitation.
Two of these factors showed significant effects. The mammoths did best with a strong seasonal variation in temperature (blue line), and one group of plants was negatively correlated with the mammoths (red).
|
The analysis shows no effect of humans on the mammoths. Not only did the 'human' data not test as having a significant effect, it is rather symmetrically distributed around zero effect.
This is the upper left part of the Extended Data Figure 4, in the online version of the article (including the pdf file).
|
The "headline story" from this work is that humans did not play a major role in the extinction of the mammoths. However, the big scientific story is the nature of the analysis, with the novel use of a large amount of environmental DNA from various places over the ages.
The headline seems simple, but the science is not. And the figure above is just a part of the analysis, chosen as an example.
The following figure hints at the complexity of the story...
The figure shows various things over the last 50,000 years.
The top set of curves shows two climate measures, which have been scaled for comparison. The blue line is for average temperature (left-hand y-axis scale); orange is for precipitation (right-hand y-axis scale).
The bottom set of curves is for the prevalence of four types of plants.
The x-axis is time, from 50,000 years ago on the left to 'now' on the right. The thick vertical gray bar is for the LGM (Last Glacial Maximum, which ended about 19,000 years ago).
Big picture: Things changed after the LGM.
This is part of Figure 2a from the article. You can see a labeled x-axis scale there. Caution... The full Figure 2a has several parts, which partially overlap. The figure shown above contains fragments of other parts at both the top and bottom.
|
Overall, the article presents a wealth of new data on biodiversity in the Arctic over the last 50,000 years. It is a time of complexity, especially after the LGM. One point that comes from this analysis is that they see little evidence for a role of humans in the extinction of the mammoths. It is more likely that the mammoths were unable to adapt to the rapidly changing food supply following the rapid melting of the glaciers.
News stories:
* Not this time: climate change, not humans, wiped out wooly mammoths -- The environment changed faster than the mammoths could adapt at the end of the last Ice Age. (Tibi Puiu, ZME Science, October 22, 2021.)
* Humans did not cause woolly mammoths to go extinct -- climate change did. New DNA research shows the world got too wet for the giant animals to survive. (Science Daily (University of Cambridge), October 20, 2021.)
The article, which is open access: Late Quaternary dynamics of Arctic biota from ancient environmental genomics. (Yucheng Wang et al, Nature 600:86, December 2, 2021.)
The second post of this set: What caused the extinction of the mammoths -- II (January 18, 2022). Immediately above.
Recent posts about mammoths:
* Briefly noted... The oldest known genome: a new record (August 31, 2021).
* Accumulation of deleterious mutations in the last mammoths (May 6, 2017). The work reported here could be related to the extinction, but, as noted in the post, there is no clear evidence for what the connection actually was.
A useful introduction to metagenomics: Is there useful ancient DNA in the dirt? (August 8, 2017).
There is more about genomes and sequencing on my page Biotechnology in the News (BITN) - DNA and the genome. It includes an extensive list of related Musings posts.
More about climate change: The effect of climate change on rainbows (January 16, 2023).
January 13, 2022
Briefly noted... Bird song from brain waves
January 13, 2022
Capture brain waves from a person, preferably using electrodes implanted in the head, and you can use them to drive motion of a limb, or get a computer to generate speech that reflects what the person intended. Do that with a song bird, and you can get the computer to generate bird song. That's the essence of an article from last summer. Why do this? Maybe scientific curiosity? However, there are parallels between sound production in song birds and humans, so this could be a useful model system.
* News story: Researchers translate a bird's brain activity into song -- Study demonstrates the possibilities of a future speech prosthesis for humans. (Science Daily (University of California - San Diego), June 18, 2021.) Links to the article.
* Direct link to the article: Neurally driven synthesis of learned, complex vocalizations. (Ezequiel M Arneodo et al, Current Biology 31:3419, August 9, 2021.) Audio files available; you may need subscription access to the article itself.
* Also see: Using a brain-computer interface to direct synthetic speech (July 16, 2019).
* My page for Biotechnology in the News (BITN) -- Other topics includes a section on Brain. It includes a list of brain-related posts.
Briefly noted... Possible detection of a planet beyond our galaxy
January 12, 2022
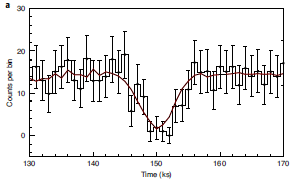
|
Graphs such as this have become familiar. The dip in intensity may be evidence that a planet has passed in front of a star. So why show this "transit" event? What's measured here is not ordinary light, but X-rays. And the star is in another galaxy. There are a lot of technical issues, but if the report holds up, it would be the best evidence yet for a planet beyond our galaxy.
This is Figure 1a from the article.
|
* News story: Astronomers Might Have Found a Planet in Another Galaxy. (Evan Gough, Universe Today, November 3, 2021. Now archived.) Links to the article.
* Direct link to the article: A possible planet candidate in an external galaxy detected through X-ray transit. (Rosanne Di Stefano et al, Nature Astronomy 5:1297, December 2021.) You may find a freely available copy by putting the article title in Google Scholar.
* A post with a similar transit observation, though not for a planet: Rings for Chariklo (May 9, 2014).
|
Fidelity of DNA replication in micro-gravity
January 11, 2022
Is traveling in space hazardous? Well, it certainly is different.
A recent article tests the effect of "micro-gravity" (i.e., weightlessness) on the accuracy of DNA replication.
The first figure summarizes the findings...
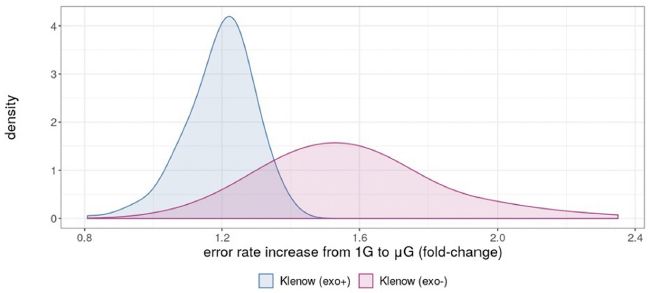
The graph shows the frequency of various error rate effects in DNA replication. The error rate effects are shown on the x-axis; the frequency is shown on the y-axis (labeled as "density", reflecting the nature of the data analysis).
For example, for one set of data (blue), the peak (most-common) error rate effect was about 1.2, according to the graph. That number is the ratio of measured error rates micro-gravity (µG) vs normal gravity (1G). That is, there was about a 20% increase in error rate in micro-gravity.
For the other data set (red), the peak ratio was near 1.6; there was about a 60% increase in the error rate during DNA replication under micro-gravity.
The results shown in this graph are a summary over many measurements, which vary widely depending on the specific base sequences being replicated. The article has extensive data for various types of mutational events.
This is Figure 3C from the article. (Caution... In the article, the parts of the figure are not labeled.)
|
What are those two data sets for? For two DNA polymerases. They are both forms of a DNA polymerase from E coli bacteria, an enzyme that is a workhorse of lab work. One form (blue) includes the proofreading part of the enzyme. The other form (red) lacks proofreading. (That proofreading part is called "exo", for exonuclease, in the labeling.)
That is, the core polymerase, without proofreading activity, gave about a 60% increase in replication errors during micro-gravity. With proofreading, it gave about a 20% increase.
Don't get bogged down in the numbers. There are a lot of data, and the effects vary with type of mutation and base sequence being replicated. The results shown here are intended to give the idea. This is the first study of DNA replication under micro-gravity. The results suggest there is a problem. That's the main point for now.
How did they do this? In an airplane, as it made a sharp change in altitude leading to a brief micro-gravity situation. This system allows measurement of the effect of the change in gravity, without interference from the radiation associated with being in space.
The following figure shows the gravity for the measurements...
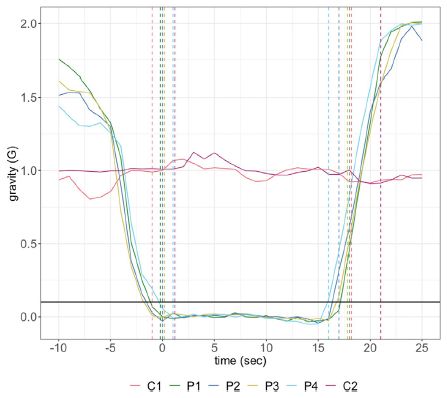
|
The graph shows gravity (y-axis) vs time (x-axis) for relevant periods in the flight.
The horizontal black line near the bottom shows 0.1 G (where G is normal gravity); that was the cut-off for determining the micro-gravity situation. You can see that the period of micro-gravity was about 20 seconds in each case.
Replication fidelity was also measured during level flight, to get data for 1 G.
This is Figure 1 from the article.
|
This article reports the effect of weightlessness (micro-gravity) on the fidelity of DNA replication, in a system independent of radiation effects. The results suggest that it is an issue that needs further attention -- especially if humans are to spend extended time in space.
News story: Cells' replication of DNA is more error-prone in microgravity. (Nanowerk News (from the journal), November 29, 2021.)
The article, which is open access: Fidelity of a Bacterial DNA Polymerase in Microgravity, a Model for Human Health in Space. (Aaron H Rosenstein & Virginia K Walker, Frontiers in Cell and Developmental Biology 9:702849, November 2021.)
Other posts about possible health issues in space:
* Improving health in space: add a little gravity (December 3, 2022).
* Which direction does blood flow in an astronaut? (January 7, 2020).
* Briefly noted... 1. The effect of space travel on humans: a study of identical twins (November 13, 2019).
A protein that can assist with handling actinium
January 9, 2022
There is interest in using radioactive actinium (element 89) for treating cancer. However, handling Ac is not easy. The amounts are minuscule, and the stuff is radioactive; half-lives of isotopes of interest are measured in days or hours.
A recent article reports a protein that binds Ac, with useful specificity. It's an interesting story.
We'll start by looking at some of the results. The first figure shows the binding of Ac, in the form of the Ac3+ ion, to the protein...
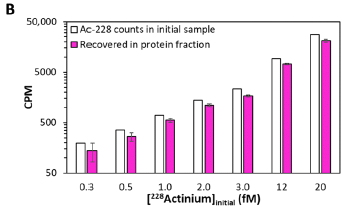
|
Each pair of bars shows the amount of Ac3+ in a solution (clear bar) and the amount from that solution that bound to a standard amount of the protein (red bar). The Ac3+ is measured as cpm (counts per minute; y-axis; log scale). The pairs of bars are for different starting concentrations of Ac3+ (x-axis).
For each case, most of the Ac3+ bound to the protein.
This is Figure 3B from the article.
|
Importantly, the binding is very strong. The lowest concentration tested here is 0.3 femtomolar (fM), and the binding seems nearly complete even at that low level. (0.3 fM Ac-228 = 7x10-14 grams per liter = 70 attograms per milliliter.)
The next figure provides some information about the specificity of the protein for binding metal ions. Caution, the figure is a bit confusing.
The top curve (red) is the one of interest. It shows the recovery of Ac3+, bound to the protein as in the previous figure. Here it is shown as a percentage -- and it is near 100% straight across.
What did they do? They added a mixture of other metal ions, ones of biological interest. They are listed just above the bottom axis: the common 2+ ions of Ca, Mg, Zn, Mn, and Cu. How much? That is shown on the x-axis, as the mole ratio of the competing ions to Ac3+. The main part of the x-axis scale ranges from 106 to over 1011. That is, the binding of Ac3+ to the protein is not affected by even a hundred billion-fold excess of common biological ions.
|
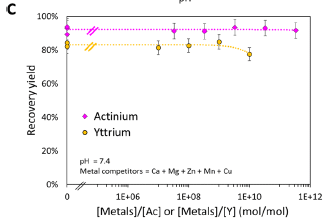
|
The figure also includes results for the binding of yttrium ions to the same protein. We'll skip that here, though the Y3+ ion is also of medical interest.
This is Figure 5C from the article.
|
It's promising. The protein could be useful in lab work, and could also be useful in the body.
Where did this protein come from? From some bacteria that, a few years ago, were found to use ions of elements in the lanthanoid group -- a novel biological finding. Investigation of the finding showed that they contained a novel protein that bound lanthanoid ions. The protein was given the name lanmodulin (by analogy to a related and well-known calcium-binding protein called calmodulin).
The lanmodulin protein was good at binding ions of the lanthanoid elements. The scientists wondered if it might also be good at binding ions of the actinoid elements, a similar group. In particular, the protein seems to have high specificity for ions with 3+ charge, a common feature in both groups.
It worked.
The current work is of interest for developing a tool for working with an unusual element, possibly useful in cancer therapy. It is also of interest for how it makes use of the new story of the biology of the lanthanoid elements. The authors also suggest that lanmodulin may be useful in studying the chemistry of other actinoid elements.
News stories:
* Radioactive metals for medicine get a boost from recently discovered protein -- Metals such as actinium could eventually be used in next-generation cancer therapies. (EurekAlert! (Penn State), October 20, 2021.)
* Cancer therapies and nuclear material detection get a boost from newly discovered protein. (Lawrence Livermore National Laboratory, October 20, 2021. Now archived.)
The article, which is open access: Capturing an elusive but critical element: Natural protein enables actinium chemistry. (Gauthier J-P Deblonde et al, Science Advances 7:eabk0273, October 20, 2021.)
More about lanmodulin: Briefly noted... A terbium detector (January 19, 2022).
Previous post about an actinoid: Einsteinium chemistry (March 9, 2021).
This post is listed on my page Introductory Chemistry Internet resources in the section Lanthanoids and actinoids.
My page for Biotechnology in the News (BITN) -- Other topics includes a section on Cancer. It includes an extensive list of relevant Musings posts.
January 7, 2022
Briefly noted... A millipede worthy of the name, at last
January 7, 2022

|
The name millipede means a thousand legs. However, the largest number of legs actually found in a "millipede" has been 750. Until now. A new article reports finding a new species of millipede with as many as 1,306 legs over 330 segments. (A defining feature of millipedes is that most body segments have two pairs of legs.) The species is named Eumillipes persephone. We should note that only one specimen out of four examined actually had >1000 legs. Still, it is progress.
|
* This is Figure 1a from the article. The specimen is about 1 millimeter wide and 100 mm long. That means legs occur every 0.15 mm.
* News story: At long last, scientists find a true millipede that has 1,000 legs (more actually). (Tibi Puiu, ZME Science, December 16, 2021.) Links to the article, which is open access.
|
Briefly noted... Does ammonia neutralize some of the sulfuric acid in the atmosphere of Venus, allowing life?
January 6, 2022
A year ago we noted an article that reported evidence for phosphine, PH3, in the atmosphere of Venus. The scientists suggested that the phosphine was evidence for life in the clouds. That article has turned out to be controversial at multiple levels, but it has also led to a stream of articles about the possibility of life in the Venusian clouds. We now have another such article -- again, intriguing and speculative. The article suggests that there is substantial ammonia, NH3, in the clouds, and that it (partially) neutralizes the sulfuric acid, at least in some places. Where does the ammonia come from? From life processes, they suggest -- lacking any good alternative. Is there any evidence for NH3 there? There actually is, though it, too, is controversial.
* As with pretty much the whole life-on-Venus story, what we really need is evidence. There are missions in the works, so over the coming years we may get evidence that allows us to evaluate the various proposals. For now, they are fun -- and speculative.
* News story: Could acid-neutralizing life-forms make habitable pockets in Venus' clouds? (Science Daily (MIT), December 20, 2021.) Links to the article, which is open access.
* Recent post about life on Venus: Briefly noted... Photosynthesis on Venus? (November 17, 2021).
* Original post on the phosphine story: Briefly noted... Phosphine on Venus? Implications for the possibility of life on Venus? (October 7, 2020). Links to follow-up posts.
Making new cartilage, using stem cells
January 5, 2022
A recent article reports progress in making cartilage in the lab.
Here is a picture of some of what they made...
|
This is part of Figure 4a from the article.
The numbers on the ruler are millimeters.
|
Here is a test of the quality of such material...
The figure shows a test of strength of natural cartilage material (left) and a sample of the new lab-grown material (right).
They are about the same. That is, the lab-grown material looks fine, as judged by this test.
ESC = embryonic stem cells. The leading h means that they are from humans.
This is Figure 2e from the article.
|
Not excited by the figures? That's ok; what's important is not easily shown, and the work is incomplete. But cartilage problems are common, and solutions are limited. The current work is about progress in making replacement cartilage.
What did they do?
The scientists started with embryonic stem cells, and differentiated them into chondrocytes (cartilage-producing cells). They then grew up these cells in the lab. After exploring various growth conditions, they were able to grow pieces a few millimeters across on a solid surface. The material generates extracellular matrix; there is no exogenous support material (commonly called "scaffold") in the tissue. Thus they made small pieces of cartilage tissue, free of extraneous material.
An interesting aspect of the work is that maintaining low oxygen levels (5% instead of the 20% typical of air) improves differentiation and development of the cartilage tissue.
What next?
The current article is about growing up pieces of cartilage. Now it is time to test whether these pieces are useful, by transplanting them into an animal.
The story here is that they have an improved process for making tissue that may be suitable for repairing cartilage. We'll see.
News stories:
* Embryonic stem cells used for cartilage regeneration. (Hannah Flynn, BioNews, October 4, 2021.)
* Power of Stem Cells Harnessed To Create Human Cartilage Tissue. (SciTechDaily (University of Southampton), October 6, 2021.)
The article, which is open access: A scaffold-free approach to cartilage tissue generation using human embryonic stem cells. (Lauren A Griffith et al, Scientific Reports 11:18921, September 28, 2021.)
Among posts about cartilage:
* Silicates and gene expression -- a new way to induce making cartilage? (June 21, 2022).
* A claim of finding dinosaur DNA (May 31, 2020).
* Humans may be more like salamanders than we had thought (limb regeneration) (February 11, 2020).
* Using your nose to fix knee damage (January 28, 2017).
* The role of zinc in arthritis (July 18, 2014).
More ESC: Synthetic mouse embryos: made without sperm, egg cell, or uterus (October 1, 2022).
There is more about stem cells and related topics, including regeneration, on my page Biotechnology in the News (BITN) for Cloning and stem cells. It includes two extensive lists of related Musings posts.
An iodine-based propulsion system tested in space
January 3, 2022
Ion-thrust engines are used in space, for tasks such as adjusting the position of satellites in orbit. Xenon gas is most commonly used as the ion source.
A recent article examines the use of iodine gas as the ion source.
The first figure compares ion formation from I2 and Xe...
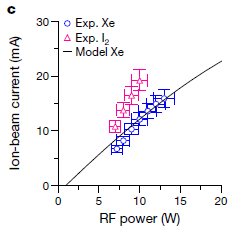
|
The graph shows ion formation, as measured by the resulting current (y-axis), vs power input (x-axis).
Results are shown for I2 gas (red) and Xe gas (blue). The results are similar. If anything, ion formation from I2 is a little more efficient.
The solid line shows what is expected for Xe; results are in good agreement with expectations.
This is Figure 2c from the article.
|
Iodine gas works for making ions.
A problem with iodine is that it is corrosive. The current work involved development of a system, based on ceramics, that was not susceptible to I2-induced corrosion.
How about a real test? The next figure shows results from a test of an I2-based thruster on a satellite in space...
The graph shows the altitude of the satellite (y-axis) over a three-month period (x-axis).
The abrupt changes were due to firing the iodine-based thruster. The first firings were to raise the satellite, to make up for the altitude it had lost during December.
The red and blue points are actual measurements. The solid line is what was intended.
The agreement is excellent.
Note that the entire range of the test was about 2 km. Individual movements of the satellite were about 200 m, took about 90 minutes, and used about 12 mg of iodine.
The test here was with a tiny CubeSat satellite, weighing about 20 kilograms. The I2-based engine weighs only a kilogram, and is about 10 cm across.
This is Figure 4a from the article.
|
This was the first test in space of an iodine-based propellant system. It worked.
Why iodine? It's cheaper than xenon -- both more abundant and easier to purify. Cost per se has typically not been a big concern for space work, though that is changing with the trend toward privatization of space. Aside from cost, iodine has an important practical advantage. Xe is a gas under ordinary conditions; storing large amounts in space requires containers that can withstand high pressures. I2 is a solid, much easier to handle and store. And it sublimates -- converts to a gas -- easily. So, when it is time to do a space maneuver, just warm up a little iodine to make the needed vapor. It works.
News stories:
* Iodine successfully tested in satellite ion thrusters. (Bob Yirka, Phys.org, November 18, 2021.)
* Iodine propulsion systems take flight in space -- Iodine-based ion propulsion could power small satellites and help solve our space junk problem. (Jessica Orwig, Inside Science, November 17, 2021. Now archived.) What is the relevance to space junk? The tiny engine could make controlled de-orbiting of small satellites practical.
* French startup demonstrates iodine propulsion in potential boost for space debris mitigation efforts. (Andrew Jones, SpaceNews, January 22, 2021.) This news story is based on the early announcement of the test, not on the article.
* News story accompanying the article: Aerospace engineering: Iodine powers low-cost engines for satellites -- Solid iodine transforms directly into gas when heated - a property that has been used to create cheap, compact engines that could make large networks of small satellites commercially viable. (Igor Levchenko & Kateryna Bazaka, Nature 599:373, November 18, 2021.)
* The article, which is open access: In-orbit demonstration of an iodine electric propulsion system. (Dmytro Rafalskyi et al, Nature 599:411, November 18, 2021.)
For background on the type of engine: Wikipedia: Ion thruster. The page includes the current article. The "comparison" table includes the iodine-based engine discussed here.
Previous post about iodine: Did the Fukushima nuclear accident lead to a burst of thyroid cancer? (July 17, 2016).
Top of page
The main page for current items is Musings.
The first archive page is Musings Archive.
E-mail announcements -- information and sign-up: e-mail announcements. The list is being used only occasionally.
Contact information
Site home page
Last update: April 6, 2026